
Individual morphometrics: the simplest GLLVM
Source:vignettes/articles/morphometrics.Rmd
morphometrics.RmdThis is the simplest case the package handles, and it is the foundation for everything else on the complexity ladder. One observational level (individuals), several continuous traits per individual, and shared low-rank covariance between traits. No phylogeny, no spatial structure, no nesting. This article walks through that model end-to-end. The working question is simple: if five body measurements are all partly driven by size and shape, can a rank-2 GLLVM recover that shared structure and still leave trait-specific variation on the diagonal?
Simulate
We pretend to measure sparrows on five body traits. The true loading matrix has a size axis (factor 1, all positive: bigger birds tend to be bigger on every measurement) and a shape axis (factor 2, mixed signs: long-and-skinny vs. short-and-stocky). Each trait also has a small trait-specific unique variance :
set.seed(2025)
n_ind <- 150
trait_names <- c("length", "mass", "wing", "tarsus", "bill")
T <- length(trait_names)
# True loadings: factor 1 = "size", factor 2 = "shape"
Lambda_true <- matrix(c(
0.9, 0.1,
1.0, 0.2,
0.8, -0.4,
0.7, -0.5,
0.6, 0.6
), nrow = T, ncol = 2, byrow = TRUE,
dimnames = list(trait_names, c("size", "shape")))
# True trait-specific unique (residual) variances
s2_true <- c(length = 0.15, mass = 0.10, wing = 0.20, tarsus = 0.12, bill = 0.18)
# Latent factors + trait-specific residuals (no global sigma_eps in truth)
u <- matrix(rnorm(n_ind * 2), n_ind, 2)
eps <- sapply(seq_len(T), function(t) rnorm(n_ind, sd = sqrt(s2_true[t])))
Y <- u %*% t(Lambda_true) + eps
rownames(Y) <- as.character(seq_len(n_ind))
colnames(Y) <- trait_names
# Wide-format data: one row per individual, one column per trait
df_wide <- data.frame(
individual = factor(seq_len(n_ind)),
Y,
check.names = FALSE
)
# Long-format data: one row per (individual, trait)
df <- data.frame(
individual = factor(rep(seq_len(n_ind), each = T)),
trait = factor(rep(trait_names, n_ind), levels = trait_names),
value = as.vector(t(Y))
)
head(df, 8)
#> individual trait value
#> 1 1 length 0.16350255
#> 2 1 mass 0.27079967
#> 3 1 wing 0.30969595
#> 4 1 tarsus 0.67902306
#> 5 1 bill -0.40893038
#> 6 2 length -0.11372224
#> 7 2 mass 0.09734399
#> 8 2 wing -0.58972821
Sigma_true <- Lambda_true %*% t(Lambda_true) + diag(s2_true) # full between-individual covariance
round(Sigma_true, 2)
#> length mass wing tarsus bill
#> length 0.97 0.92 0.68 0.58 0.60
#> mass 0.92 1.14 0.72 0.60 0.72
#> wing 0.68 0.72 1.00 0.76 0.24
#> tarsus 0.58 0.60 0.76 0.86 0.12
#> bill 0.60 0.72 0.24 0.12 0.90Pitfall: factor level order. The line
factor(rep(trait_names, n_ind), levels = trait_names)is critical. Without the explicitlevels = trait_names, R sorts levels alphabetically (bill, length, mass, tarsus, wing), and the rows of fitted will not match the rows of yourLambda_true. The model is correct; the comparison is not. This is a subtle data-prep gotcha worth flagging because it makes “recovery” look broken when only the row labels are misaligned.
Fit
The same model can be fitted from long or wide data. The long form is
canonical because it makes the (individual, trait) rows
explicit; the wide forms are the convenient equivalents for readers who
already have one row per individual and one column per trait.
# Long format (canonical)
fit_long <- gllvmTMB(
value ~ 0 + trait +
latent(0 + trait | individual, d = 2) +
unique(0 + trait | individual),
data = df,
unit = "individual" # tell the engine which column groups observations
)
# Wide data-frame format (same model with compact formula syntax)
fit_wide_formula <- gllvmTMB(
traits(length, mass, wing, tarsus, bill) ~ 1 +
latent(1 | individual, d = 2) +
unique(1 | individual),
data = df_wide,
unit = "individual"
)
# Wide matrix format (same model for matrix-first workflows)
fit_wide_matrix <- gllvmTMB_wide(Y, d = 2, family = gaussian())
# The rest of the article uses the canonical long-format fit.
fit <- fit_long
fit
#> Stacked-trait gllvmTMB fit
#> Traits = 5, individual = 150
#> Covstructs: latent_unit, unique_unit
#> Fixed effects (b_fix): 5
#> log L = -714.850 convergence = 0
#> Note: Lambda_B identified up to rotation (use suggest_lambda_constraint() or rotate_loadings()).
#> Run gllvmTMB_diagnose(fit) for a full health check, or summary(fit) for parameter estimates.
cat("Long/formula-wide logLik difference:",
signif(abs(as.numeric(logLik(fit_long)) -
as.numeric(logLik(fit_wide_formula))), 3),
"\n")
#> Long/formula-wide logLik difference: 0
cat("Long/matrix-wide logLik difference:",
signif(abs(as.numeric(logLik(fit_long)) -
as.numeric(logLik(fit_wide_matrix))), 3),
"\n")
#> Long/matrix-wide logLik difference: 1.02e-08gllvmTMB defaults to unit = "site", meaning it looks for
a column literally named site in the data. This
morphometric data set has no site column — the grouping
factor is individual. Passing
unit = "individual" redirects the engine to the correct
column; everything else is standard GLLVM machinery.
traits(length, mass, wing, tarsus, bill) is the
formula-level wide data-frame marker. Because the response traits are
named on the LHS, the RHS can use the compact shorthand: 1
becomes the trait-specific intercepts 0 + trait, and
latent(1 | individual) becomes
latent(0 + trait | individual).
gllvmTMB_wide(Y, ...) is the matrix-in entry point for the
same model when the response already lives in a numeric matrix.
What model did we fit?
For each individual we observe a vector of trait measurements — for example, body length, mass, wing chord, tarsus, bill depth in a sparrow study; tail length, fin span, gill area in a fish study; or six items on a psychometric scale in a behavioural study. The reduced-rank GLLVM is
where
is the loading vector for trait
,
is a
-dimensional
latent factor for individual
,
and
is the trait-specific residual with variance
.
The unique() term collects those residual variances on the
diagonal of
.
The implied between-individual trait covariance is
For traits and factors this is still a small example; the practical gain is interpretability, not a dramatic parameter saving. As grows, the low-rank part scales with loadings rather than the full covariance surface. That low-rank structure is the point of GLLVMs: shared biological axes first, trait-specific variation second.
+ unique(0 + trait | individual)with cross-sectional data. With cross-sectional data (one observation per individual per trait), the(trait, individual)pair uniquely identifies every row. gllvmTMB detects this automatically: it fixes to a negligibly small constant (and prints ani Auto-suppressing sigma_eps: ...message), so theunique()term absorbs the full observation-level residual and becomes the trait-specific residual variance . The confounding that would arise if both and were estimated freely is therefore resolved by construction — you do not need to choose between them. With longitudinal data (multiple sessions per individual), both terms are separately identified and can be estimated without any auto-suppression; theunique()term then captures genuine between-individual trait-specific variance on top of the latent structure. See the Covariance & correlation article for the side-by-side comparison.
Recovered vs truth
Sigma_hat <- extract_Sigma(fit, level = "unit")$Sigma
round(Sigma_hat, 2)
#> length mass wing tarsus bill
#> length 0.94 0.91 0.73 0.50 0.69
#> mass 0.91 1.16 0.83 0.57 0.80
#> wing 0.73 0.83 1.11 0.73 0.40
#> tarsus 0.50 0.57 0.73 0.72 0.17
#> bill 0.69 0.80 0.40 0.17 1.01
cat("Frobenius |Sigma_hat - Sigma_true| / |Sigma_true| =",
round(norm(Sigma_hat - Sigma_true, "F") / norm(Sigma_true, "F"), 3), "\n")
#> Frobenius |Sigma_hat - Sigma_true| / |Sigma_true| = 0.117A side-by-side correlation heatmap shows whether the recovered trait correlations agree with truth at the entry level:
suppressPackageStartupMessages(library(ggplot2))
make_corr_long <- function(M, label) {
R <- cov2cor(M)
data.frame(
trait_i = factor(rep(rownames(R), ncol(R)), levels = rownames(R)),
trait_j = factor(rep(colnames(R), each = nrow(R)), levels = colnames(R)),
rho = as.vector(R),
panel = label
)
}
rownames(Sigma_true) <- colnames(Sigma_true) <- trait_names
rownames(Sigma_hat) <- colnames(Sigma_hat) <- trait_names
df_corr <- rbind(make_corr_long(Sigma_true, "True"),
make_corr_long(Sigma_hat, "Fitted"))
df_corr$panel <- factor(df_corr$panel, levels = c("True", "Fitted"))
ggplot(df_corr, aes(trait_j, trait_i, fill = rho)) +
geom_tile(colour = "white") +
geom_text(aes(label = sprintf("%.2f", rho)), size = 3.2) +
scale_fill_gradient2(low = "#3b82c6", mid = "white", high = "#d33",
midpoint = 0, limits = c(-1, 1)) +
facet_wrap(~ panel) +
scale_y_discrete(limits = rev) +
labs(x = NULL, y = NULL, fill = expression(rho)) +
theme_minimal(base_size = 11) +
theme(panel.grid = element_blank(),
strip.text = element_text(face = "bold"))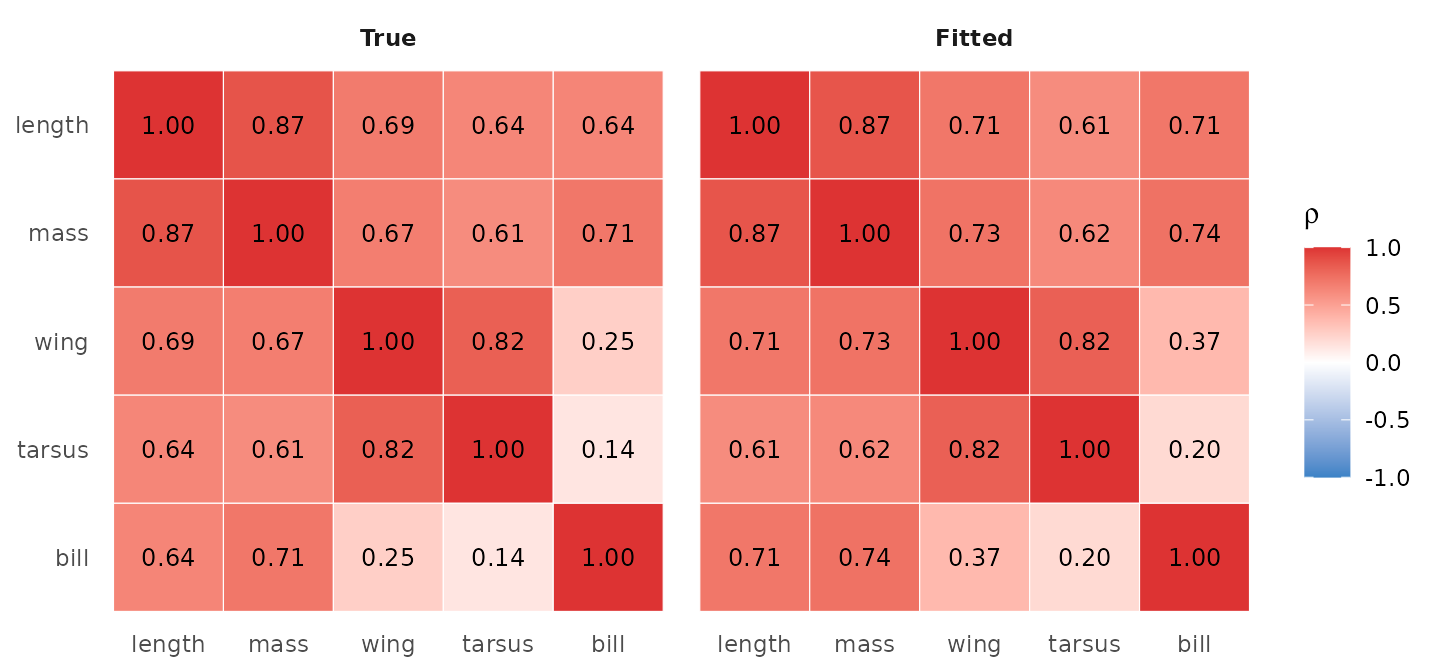
True (left) vs fitted (right) trait-pair correlations from Sigma_B = Lambda Lambda^T. The 1:1 agreement of the colours and labels is the visual signature of clean recovery.
What
extract_Sigma(level = "unit")$Sigmareturns. The total between-unit covariance — the sum of the reduced-rank shared component and the diagonal unique component. If you fit without+ unique(), and you get only , which understates the diagonal. Adding+ unique()(as in the fit above) fills in the diagonal correctly. Usepart = "shared"/part = "unique"to pull just one component.
Pairwise correlations and communality
For inference on individual cross-trait correlations or per-trait
communality, the canonical extractors return tidy data frames.
extract_correlations() defaults to
method = "fisher-z" (fast Fisher-transform Wald CI; bounds
guaranteed in
).
Communality point estimates are enough for this first worked example.
When interval estimates matter, prefer profile or Wald-style intervals
where the target supports them. Bootstrap is a slower, deliberate option
for a separate cached check, not part of this rendered article.
extract_correlations(fit, tier = "unit")
#> tier trait_i trait_j correlation lower upper method
#> 1 B length mass 0.8711314 0.8262171 0.9050412 fisher-z
#> 2 B length wing 0.7124695 0.6233873 0.7832935 fisher-z
#> 3 B mass wing 0.7308546 0.6462919 0.7976850 fisher-z
#> 4 B length tarsus 0.6068548 0.4947062 0.6991228 fisher-z
#> 5 B mass tarsus 0.6203449 0.5108731 0.7100183 fisher-z
#> 6 B wing tarsus 0.8188765 0.7581044 0.8655483 fisher-z
#> 7 B length bill 0.7106087 0.6210775 0.7818326 fisher-z
#> 8 B mass bill 0.7372131 0.6542491 0.8026447 fisher-z
#> 9 B wing bill 0.3730787 0.2263507 0.5032508 fisher-z
#> 10 B tarsus bill 0.1979495 0.0389227 0.3471966 fisher-z
extract_communality(fit, level = "unit")
#> length mass wing tarsus bill
#> 0.8444800 0.8986996 0.8233290 0.8598782 0.8286985Communality is the fraction of trait ’s variance shared with the others via the latent factors; high means strongly integrated, low means trait-specific variation dominates.
Recovery across replicates
A single fit cannot tell you whether the model is recovering truth or
just one lucky draw. Repeat over R = 20 independently
simulated datasets and look at the distribution of the bias. The loop
uses the same long/wide formula pair as above; it checks the wide
equivalent on the first replicate only, so the article does not double
the cost of every recovery fit.
run_one <- function(seed) {
set.seed(seed)
u <- matrix(rnorm(n_ind * 2), n_ind, 2)
eps <- sapply(seq_len(T), function(t) rnorm(n_ind, sd = sqrt(s2_true[t])))
Y <- u %*% t(Lambda_true) + eps
colnames(Y) <- trait_names
df_rep_wide <- data.frame(
individual = factor(seq_len(n_ind)),
Y,
check.names = FALSE
)
df_rep <- data.frame(
individual = factor(rep(seq_len(n_ind), each = T)),
trait = factor(rep(trait_names, n_ind), levels = trait_names),
value = as.vector(t(Y))
)
fit_rep_long <- suppressMessages(suppressWarnings(gllvmTMB(
value ~ 0 + trait +
latent(0 + trait | individual, d = 2) +
unique(0 + trait | individual),
data = df_rep, unit = "individual"
)))
if (identical(seed, 1L)) {
fit_rep_wide <- suppressMessages(suppressWarnings(gllvmTMB(
traits(length, mass, wing, tarsus, bill) ~ 1 +
latent(1 | individual, d = 2) +
unique(1 | individual),
data = df_rep_wide, unit = "individual"
)))
stopifnot(
abs(as.numeric(logLik(fit_rep_long)) -
as.numeric(logLik(fit_rep_wide))) < 1e-6
)
}
Sigma_hat_rep <- extract_Sigma(fit_rep_long, level = "unit")$Sigma
data.frame(
seed = seed,
entry = paste0("[", rep(1:T, each = T), ",", rep(1:T, T), "]"),
true = as.vector(Sigma_true),
fitted = as.vector(Sigma_hat_rep)
)
}
rep_df <- do.call(rbind, lapply(1:20, run_one))
ggplot(rep_df, aes(true, fitted)) +
geom_abline(slope = 1, intercept = 0, colour = "grey40", linetype = 2) +
geom_point(alpha = 0.4, colour = "#1f78b4", size = 1.6) +
labs(x = expression("True"~Sigma[B]~"entry"),
y = expression("Fitted"~hat(Sigma)[B]~"entry"),
title = "Recovery on 20 replicate datasets, n = 150 individuals") +
theme_minimal(base_size = 11)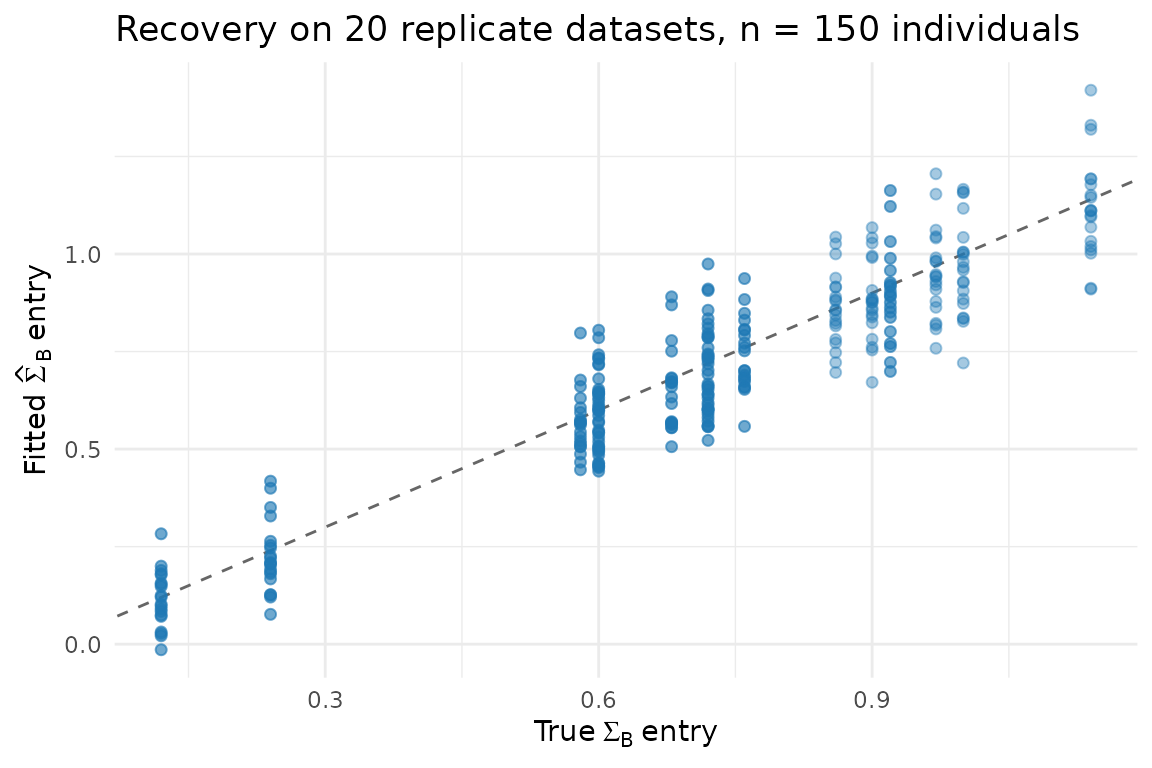
rep_df |>
mutate(off_diag = sub("\\[(.),(.)\\]", "\\1\\2", entry),
off_diag = substr(off_diag, 1, 1) != substr(off_diag, 2, 2)) |>
group_by(off_diag) |>
summarise(
bias = mean(fitted - true),
rmse = sqrt(mean((fitted - true)^2)),
.groups = "drop"
)
#> # A tibble: 2 × 3
#> off_diag bias rmse
#> <lgl> <dbl> <dbl>
#> 1 FALSE -0.0184 0.113
#> 2 TRUE -0.0169 0.0990The off-diagonal entries (the cross-trait covariances — the biology) are recovered with bias close to zero. Diagonal entries are slightly under at , which is the expected finite-sample behaviour and shrinks as grows.
Why latent + unique? dep and
indep on the same data
The morphometric workflow pairs latent(d = K) with
unique() to estimate
.
Two alternative covstruct modes — dep (full unstructured
with
free entries) and indep (diagonal-only
,
no cross-trait modelling) — are also valid keywords. Refit the same data
under each to see what the user actually loses or gains.
fit_full_long <- gllvmTMB(
value ~ 0 + trait +
latent(0 + trait | individual, d = T),
data = df, unit = "individual"
)
fit_full_wide <- gllvmTMB(
traits(length, mass, wing, tarsus, bill) ~ 1 +
latent(1 | individual, d = T),
data = df_wide, unit = "individual"
)
fit_dep_long <- gllvmTMB(
value ~ 0 + trait +
dep(0 + trait | individual),
data = df, unit = "individual"
)
fit_dep_wide <- gllvmTMB(
traits(length, mass, wing, tarsus, bill) ~ 1 +
dep(1 | individual),
data = df_wide, unit = "individual"
)
fit_indep_long <- gllvmTMB(
value ~ 0 + trait +
indep(0 + trait | individual),
data = df, unit = "individual"
)
fit_indep_wide <- gllvmTMB(
traits(length, mass, wing, tarsus, bill) ~ 1 +
indep(1 | individual),
data = df_wide, unit = "individual"
)
fit_full <- fit_full_long
fit_dep <- fit_dep_long
fit_indep <- fit_indep_long
cat("Max long/wide logLik difference across comparison fits:",
signif(max(abs(c(
as.numeric(logLik(fit_full_long)) - as.numeric(logLik(fit_full_wide)),
as.numeric(logLik(fit_dep_long)) - as.numeric(logLik(fit_dep_wide)),
as.numeric(logLik(fit_indep_long)) - as.numeric(logLik(fit_indep_wide))
))), 3),
"\n")
#> Max long/wide logLik difference across comparison fits: 0
S1 <- extract_Sigma(fit, level = "unit")$Sigma
S2 <- extract_Sigma(fit_full, level = "unit")$Sigma
S3 <- extract_Sigma(fit_dep, level = "unit")$Sigma
S4 <- extract_Sigma(fit_indep, level = "unit")$Sigma
off <- function(M) M[upper.tri(M)]
truth_off <- off(Sigma_true)
ll <- function(f) round(as.numeric(logLik(f)), 2)
data.frame(
mode = c("latent(d=2) + unique", "latent(d=T) only",
"dep", "indep"),
logL = c(ll(fit), ll(fit_full), ll(fit_dep), ll(fit_indep)),
off_diag_corr_truth = c(round(cor(off(S1), truth_off), 3),
round(cor(off(S2), truth_off), 3),
round(cor(off(S3), truth_off), 3),
NA),
diag_min_max = c(sprintf("%.2f - %.2f", min(diag(S1)), max(diag(S1))),
sprintf("%.2f - %.2f", min(diag(S2)), max(diag(S2))),
sprintf("%.2f - %.2f", min(diag(S3)), max(diag(S3))),
sprintf("%.2f - %.2f", min(diag(S4)), max(diag(S4))))
)
#> mode logL off_diag_corr_truth diag_min_max
#> 1 latent(d=2) + unique -714.85 0.952 0.72 - 1.16
#> 2 latent(d=T) only -714.80 0.952 0.65 - 1.09
#> 3 dep -714.80 0.952 0.65 - 1.09
#> 4 indep -1055.50 NA 0.72 - 1.16What this shows:
-
depis mathematically identical tolatent(d = T)standalone — same logL, same to numerical precision. The two keywords are aliases of the saturated unstructured fit. -
Reduced-rank
latent(d = 2) + uniquematches the saturated fit within 0.1 nats of logL. With traits and a rank-2 truth, the rank constraint loses essentially no information and gives the same off-diagonal correlation pattern as the saturated fit ( vs truth ≈ 0.95 for both). -
indeprecovers the diagonals fine — Gaussian gives per-trait identifiability for the marginal variance even with no cross-trait modelling — but loses every off-diagonal. The off-diagonal correlation with truth isNAbecause the fitted off-diagonals are exactly zero by construction. logL is materially worse.
This is the Gaussian counterpart to the binary
comparison in Joint species distribution.
The qualitative ordering is the same — latent + unique ≥
dep ≫ indep in modelling flexibility — but the
quantitative gap between indep and the others is much
smaller for Gaussian (a few hundred nats here, vs indep
being pinned at zero diagonals for binary 1-obs-per-cell). Pick
latent + unique when you have a strong
rank-
hypothesis (size + shape, here); fall back to dep if you
suspect
and want the saturated fit; reach for indep only when you
genuinely believe traits are mutually independent — a rare default for
ecological / morphological data.
What the latent factors mean
Every factor model has a rotational ambiguity: and produce the same for any orthogonal . To compare loadings between fits or between packages, rotate to a standard convention (varimax for interpretability, or lower-triangular for a unique fit):
L_hat <- extract_ordination(fit, level = "unit")$loadings
rot <- rotate_loadings(fit, level = "unit", method = "varimax")
round(rot$Lambda, 2)
#> [,1] [,2]
#> length 0.72 0.53
#> mass 0.83 0.60
#> wing 0.36 0.89
#> tarsus 0.12 0.78
#> bill 0.91 0.08
ord <- extract_ordination(fit, level = "unit")
head(ord$scores)
#> LV1 LV2
#> 1 0.2459334 0.8620479
#> 2 -0.1122392 0.4233450
#> 3 0.6767902 0.6307250
#> 4 1.3933275 1.8019347
#> 5 0.4073921 0.3723656
#> 6 0.1376152 -0.1679856
plot(fit, type = "ordination", level = "unit")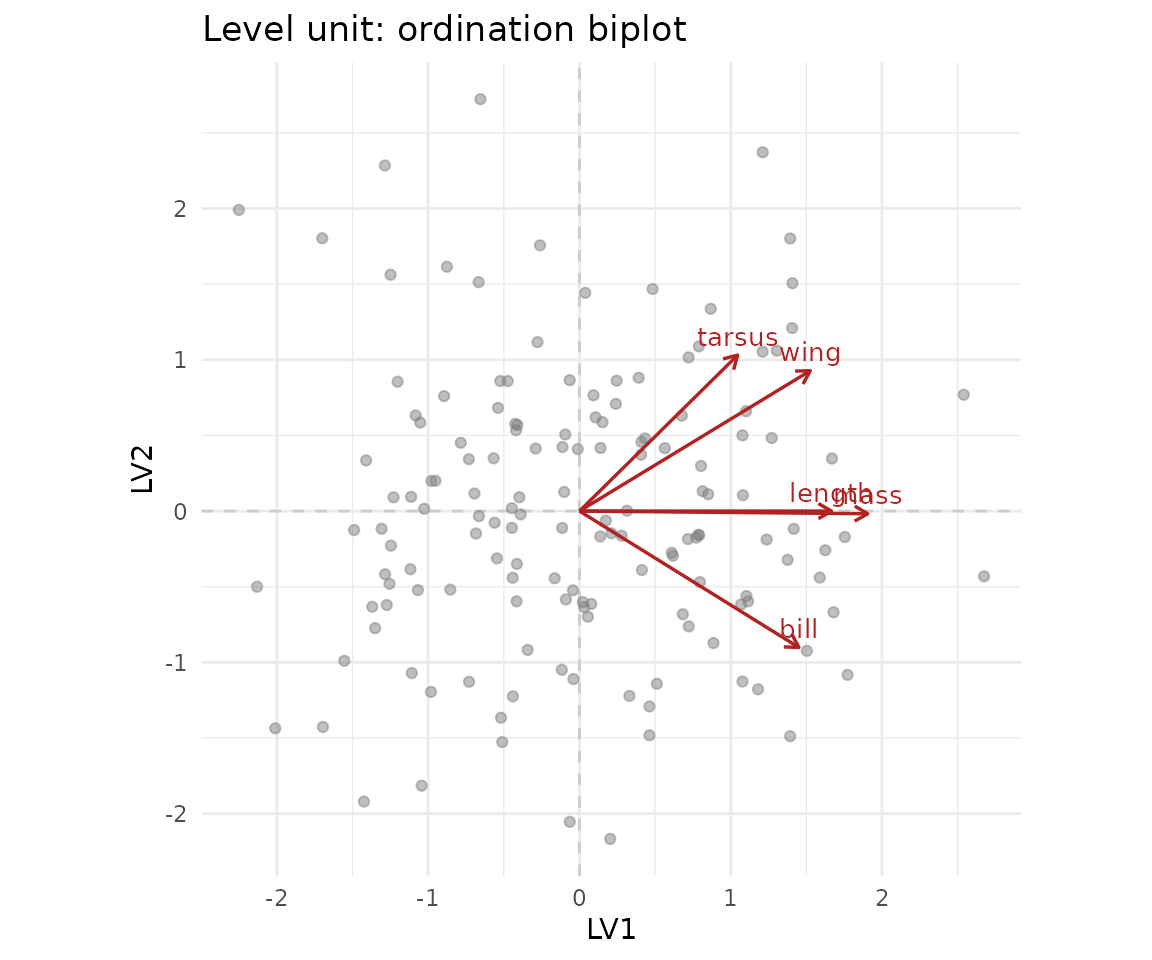
The biplot shows the trait loadings on the two latent axes. In a real morphometric study, the first axis usually picks up overall size, with all traits loading positively — the famous “general factor” of body size. The second axis picks up shape contrasts.
What this article does not cover
This is the bottom rung of the ladder. The next rungs add structure:
| Rung | New piece | Article |
|---|---|---|
| 1 | Discrete (binary) responses, individuals as sites | Joint species distribution |
| 2 | Phylogenetic dependence between rows | Phylogenetic GLLVM |
| 3 | Spatial dependence between rows | SPDE benchmark |
| 4 | Two observational levels (between- and within-unit) | Two-stage workflow |
| 5 | All of the above at once | Functional biogeography |
Read the rungs in order if you are new to GLLVMs. Jump straight to the rung that matches your data if you are not.
See also
- Simulation-based recovery study — the same experiment as the “Recovery across replicates” section above, but extended to every level on the ladder.
-
Cross-package validation
— log-likelihood agreement against
glmmTMB::rr()on identical data. - Lambda constraints — pin specific entries of when you have prior knowledge of the factor structure (for example, a confirmatory factor analysis).