
Covariance and correlation: when you need `unique()`
Source:vignettes/articles/covariance-correlation.Rmd
covariance-correlation.RmdSuppose you want between-individual correlations among behaviours
such as boldness, exploration, activity, aggression, and sociability. If
you fit only latent(), the recovered correlations can be
too large because the model has no trait-specific variance term. Pair
latent() with unique() when trait-specific
variance should stay in the denominator of the correlation matrix.
fit_latent_only <- gllvmTMB(
value ~ 0 + trait + latent(0 + trait | individual, d = 2),
data = df, unit = "individual"
)
fit_latent_unique <- gllvmTMB(
value ~ 0 + trait +
latent(0 + trait | individual, d = 2) +
unique(0 + trait | individual),
data = df, unit = "individual"
)If your data are wide, with one row per individual and one column per
behaviour, use the same gllvmTMB() entry point with
traits(...) on the left-hand side:
fit_latent_unique <- gllvmTMB(
traits(boldness, exploration, activity, aggression, sociability) ~
1 + latent(1 | individual, d = 2) + unique(1 | individual),
data = df_wide, unit = "individual"
)The decomposition
For the latent() + unique() model, the implied trait
covariance is
where level names the covariance tier being extracted,
such as unit for between-individual covariance or
unit_obs for within-individual or observation-level
covariance.
is the
loading matrix (from the latent() term) and
is a
diagonal matrix of trait-specific unique variances (from the
unique() term). The correlation matrix is then
Both pieces matter: the diagonal of is — the sum of squared loadings plus the trait-specific unique variance. Drop and the diagonal shrinks; the off-diagonals stay the same; swells.
A side-by-side demonstration
Simulate one Gaussian dataset where the true between-individual covariance has both a shared low-rank component and a trait-specific unique diagonal, then fit it two ways:
-
Model A:latent(0 + trait | individual, d = 2)only -
Model B:latent(0 + trait | individual, d = 2) + unique(0 + trait | individual)
set.seed(2026)
n_ind <- 200
trait_names <- c("boldness", "exploration", "activity",
"aggression", "sociability")
T <- length(trait_names)
## True between-individual covariance Sigma_unit = Lambda Lambda^T + diag(s)
Lambda_true <- matrix(c(
0.9, 0.1,
1.0, 0.2,
0.8, -0.4,
0.7, -0.5,
0.6, 0.6
), nrow = T, ncol = 2, byrow = TRUE,
dimnames = list(trait_names, c("syndrome_1", "syndrome_2")))
S_true <- c(0.30, 0.20, 0.40, 0.25, 0.35)
Sigma_true <- Lambda_true %*% t(Lambda_true) + diag(S_true)
sigma2_eps_true <- 0.05 # tiny observation-level noise
u <- matrix(rnorm(n_ind * 2), n_ind, 2)
e <- matrix(rnorm(n_ind * T, sd = sqrt(S_true[col(matrix(0, n_ind, T))])),
n_ind, T)
y <- u %*% t(Lambda_true) + e +
matrix(rnorm(n_ind * T, sd = sqrt(sigma2_eps_true)), n_ind, T)
df <- data.frame(
individual = factor(rep(seq_len(n_ind), each = T)),
trait = factor(rep(trait_names, n_ind), levels = trait_names),
value = as.vector(t(y))
)
fit_A <- gllvmTMB(
value ~ 0 + trait + latent(0 + trait | individual, d = 2),
data = df, unit = "individual"
)
fit_B <- gllvmTMB(
value ~ 0 + trait +
latent(0 + trait | individual, d = 2) +
unique(0 + trait | individual),
data = df, unit = "individual"
)Now extract the implied and between-individual correlation matrix from each fit:
ext_A <- suppressMessages(
extract_Sigma(fit_A, level = "unit", part = "total")
)
ext_B <- extract_Sigma(fit_B, level = "unit", part = "total")Notice that the call on fit_A emits a one-shot
advisory explaining that without unique()
the returned
is latent-only (we suppressed the message above for clean output but the
message is also stored in ext_A$note).
ext_A$note # advisory: diag missing
#> [1] "Sigma_unit is currently latent-only (Lambda Lambda^T) because no `unique(0 + trait | individual)` term is in the formula. Trait-specific unique variance is not modelled, so correlations from this matrix overstate cross-trait coupling. For the correct decomposition Sigma = Lambda Lambda^T + S, refit with `+ unique(0 + trait | individual)`."The correlations side by side
make_long <- function(R, label) {
data.frame(
trait_i = factor(rep(rownames(R), ncol(R)), levels = rownames(R)),
trait_j = factor(rep(colnames(R), each = nrow(R)), levels = colnames(R)),
rho = as.vector(R),
panel = label
)
}
R_true_named <- cov2cor(Sigma_true)
rownames(R_true_named) <- colnames(R_true_named) <- trait_names
df_corr <- rbind(
make_long(R_true_named, "1. Truth"),
make_long(ext_A$R, "2. Model A (latent only)"),
make_long(ext_B$R, "3. Model B (latent + unique)")
)
df_corr$panel <- factor(df_corr$panel, levels = c(
"1. Truth", "2. Model A (latent only)", "3. Model B (latent + unique)"))
ggplot(df_corr, aes(trait_j, trait_i, fill = rho)) +
geom_tile(colour = "white") +
geom_text(aes(label = sprintf("%.2f", rho)), size = 3) +
scale_fill_gradient2(low = "#3b82c6", mid = "white", high = "#d33",
midpoint = 0, limits = c(-1, 1)) +
scale_y_discrete(limits = rev) +
facet_wrap(~ panel) +
labs(x = NULL, y = NULL, fill = expression(rho)) +
theme_minimal(base_size = 10) +
theme(panel.grid = element_blank(),
strip.text = element_text(face = "bold"),
axis.text.x = element_text(angle = 35, hjust = 1))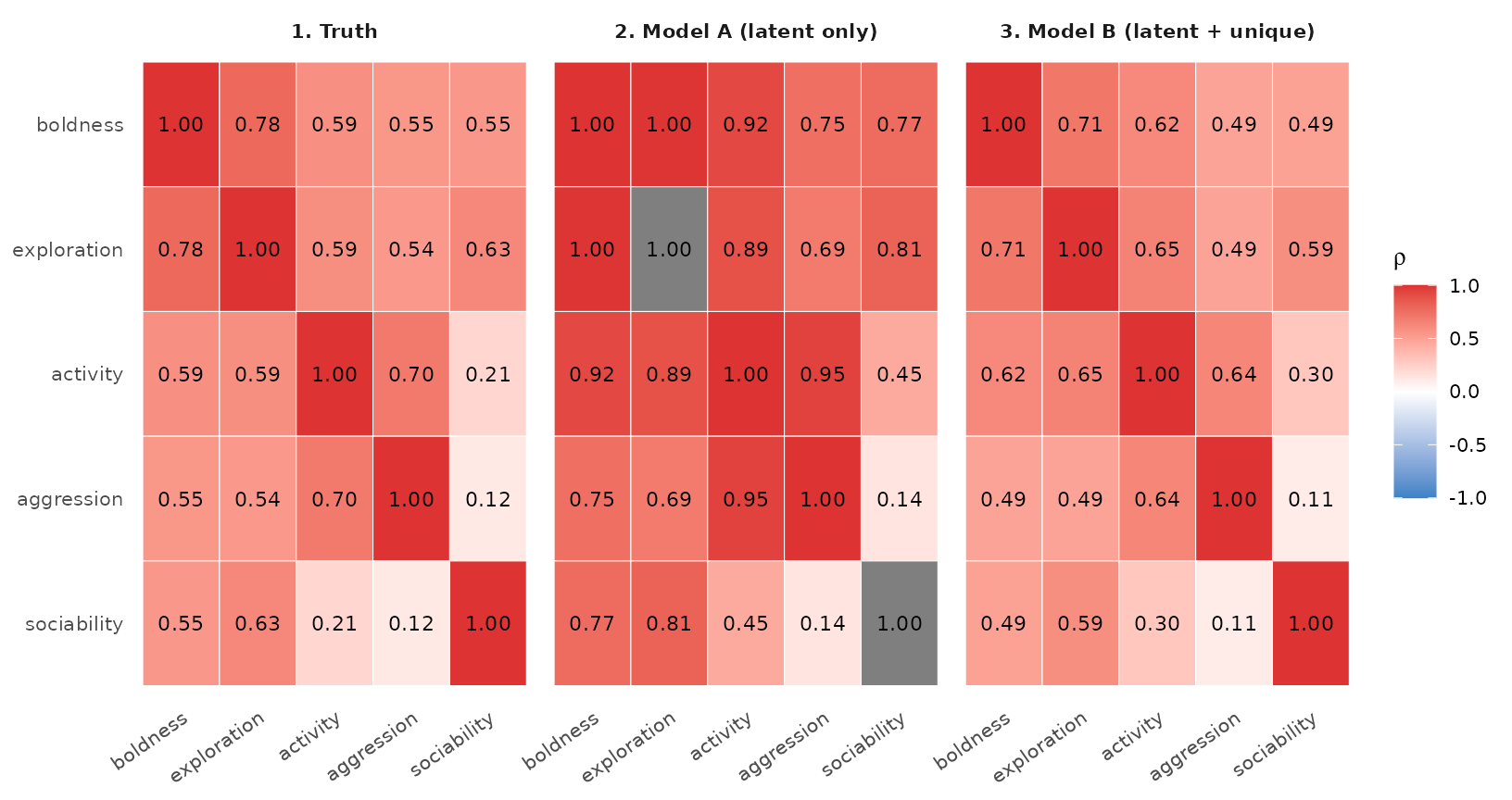
True (left) vs. recovered correlations from Model A (latent only) and Model B (latent + unique). Model A’s off-diagonals are all systematically larger than truth — that’s the inflation unique() prevents.
The middle panel (Model A) shows correlations that are
uniformly larger than truth (left panel). The right
panel (Model B) recovers the true correlations to within
sampling noise. The mechanism: Model A’s
has small diagonals (only
),
so when we divide by
the off-diagonals get inflated. Model B’s
has the full
— diagonals at the right scale and correlations at the right
magnitude.
What extract_Sigma() gives you
The unified extractor exposes the three parts of the decomposition explicitly:
# Lambda Lambda^T alone (the "shared" component)
extract_Sigma(fit_B, level = "unit", part = "shared")$Sigma |> round(2)
#> boldness exploration activity aggression sociability
#> boldness 0.71 0.79 0.67 0.52 0.52
#> exploration 0.79 0.90 0.71 0.53 0.63
#> activity 0.67 0.71 0.75 0.67 0.31
#> aggression 0.52 0.53 0.67 0.64 0.11
#> sociability 0.52 0.63 0.31 0.11 0.65
# diag(s_unit) — trait-specific unique variances (the "unique" component)
extract_Sigma(fit_B, level = "unit", part = "unique")$s |> round(2)
#> boldness exploration activity aggression sociability
#> 0.38 0.24 0.31 0.38 0.36
# Sigma_unit = Lambda Lambda^T + diag(s_unit) (the "total" — what you usually want)
extract_Sigma(fit_B, level = "unit", part = "total")$Sigma |> round(2)
#> boldness exploration activity aggression sociability
#> boldness 1.09 0.79 0.67 0.52 0.52
#> exploration 0.79 1.13 0.71 0.53 0.63
#> activity 0.67 0.71 1.07 0.67 0.31
#> aggression 0.52 0.53 0.67 1.02 0.11
#> sociability 0.52 0.63 0.31 0.11 1.01part = "total" is the default and is what you almost
always want for reporting. "shared" is useful if you
specifically want the latent-implied component (e.g. for ordination
interpretation or communality). "unique" gives the diagonal
of
as a named numeric vector.
Communality and ICC need the full too
Two trait-level summaries used in behavioural-syndromes and phenotypic-integration workflows both depend on the full decomposition. If you compute them from a latent-only fit you get the wrong answer.
Communality
This is the proportion of trait ’s variance that is shared with the other traits via the latent factors. It is bounded between 0 and 1 and is a proportion-of-variance summary, analogous in scale to (heritability) or .
The catch: with latent-only fits,
for every trait by construction, so
for every trait — the communality is identically 1 and tells you
nothing. Adding unique() gives the denominator a per-trait
slot, and communality becomes informative.
extract_communality(fit_A, level = "unit") # latent-only fit: all = 1, useless
#> boldness exploration activity aggression sociability
#> 1 1 1 1 1
extract_communality(fit_B, level = "unit") # latent + unique: meaningful
#> boldness exploration activity aggression sociability
#> 0.6481024 0.7914554 0.7068777 0.6264135 0.6440561Site-level / individual-level ICC
For two-level fits the site-level (individual-level) ICC is
i.e. the proportion of total trait variance attributable to between-
unit differences. Each piece needs the full decomposition for its level
—
and
.
Both unique() terms are required (one per level) for
to be honest.
In gllvmTMB this is
extract_ICC_site(fit)— see the recommended two-unique()
pattern below.
When unique() is not the right term
Three situations need care:
1. Binary responses (binomial family)
For binary outcomes the latent-scale residual is fixed by the link function:
| Link | Implicit residual variance |
|---|---|
logit |
|
probit |
|
cloglog |
This implicit residual plays the role of
on the latent scale. Adding an explicit unique() term on
top of binary data is typically not identified — the engine cannot
distinguish between unique() variance and the link’s
implicit residual.
extract_Sigma() has a
link_residual = "auto" option that adds the link-specific
implicit residual to
for the marginal latent-scale interpretation:
# (illustrative — not run; needs a binary fit)
extract_Sigma(fit_binary, level = "unit", part = "total",
link_residual = "auto")2. Phylogenetic tiers
phylo_latent(species, d = K) gives the shared
phylogenetic component
.
If you also need trait-specific phylogenetic variance, the companion is
phylo_unique(species), not the ordinary non-phylogenetic
unique(0 + trait | species) term. The ordinary
unique() term belongs to the non-phylogenetic species
tier.
With both phylogenetic and non-phylogenetic species components, the decomposition is
extract_Sigma(fit, level = "phy", part = "shared")
returns the phylogenetic shared component; part = "unique"
returns the
diagonal when phylo_unique() is fitted; and
part = "total" returns
.
When unit = "species",
extract_Sigma(fit, level = "unit") returns
.
Their sum is
;
extract_Omega(fit) is the convenience extractor for that
total.
Two-level (between + within) models: two unique()
terms
For repeated-measures data the recommended pattern is two
latent() + unique() pairs (Nakagawa et al. in
prep):
fit_two_level <- gllvmTMB(
value ~ 0 + trait +
latent(0 + trait | individual, d = d_B) +
unique(0 + trait | individual) +
latent(0 + trait | obs_id, d = d_W) +
unique(0 + trait | obs_id),
data = df,
unit = "individual",
unit_obs = "obs_id"
)giving
(behavioural syndromes — between-individual covariance) and
(integrated plasticity — within-individual covariance). Each
level has its own unique().
Observation-level random effects for non-Gaussian fits
For overdispersed Poisson and similar count families, the latent-scale “unique component” is recovered by adding an observation-level random effect (OLRE; Nakagawa & Schielzeth 2010):
The
on the latent log scale plays the role of
.
Native OLRE support is available: add an obs_id column (one
level per row), include unique(0 + trait | obs_id) in the
formula, pass unit_obs = "obs_id", and use
extract_residual_split(fit) to separate
(the estimated OLRE variance) from
(the distribution-specific latent residual; Nakagawa & Schielzeth
2010; Nakagawa, Johnson & Schielzeth 2017).
Summary
| You want… | You need | Notes |
|---|---|---|
| Cross-trait correlations on a Gaussian / lognormal / Gamma fit |
latent() + unique() at every level |
Otherwise correlations are inflated. |
| Correlations on a binary fit |
latent() only; link_residual = "auto" for
marginal scale |
Implicit residual depends on link (π²/3, 1, π²/6). |
| Phylogenetic decomposition |
phylo_latent() + phylo_unique() for the phylogenetic
tier; add latent() + unique() for the non-phylogenetic
species tier |
,
with each
using shared + diag(s) pieces. |
| Communality |
extract_communality(fit) + the right latent/unique
pattern |
Communality formula uses Σ; needs full Σ to be meaningful. |
Lambda Lambda^T only |
extract_Sigma(fit, part = "shared") |
Useful for ordination. |
S only |
extract_Sigma(fit, part = "unique") |
Trait-specific unique variances as a vector. |
See also
- Pitfalls — common mistakes including correlation inflation
-
?unique— the formula keyword for the unique diagonal -
?extract_Sigma— the unified extractor -
?suggest_lambda_constraint— a helper for confirmatory loading structures - Choose your model — decision tree
- Nakagawa & Schielzeth (2010) Repeatability for Gaussian and non-Gaussian data: a practical guide for biologists. Biological Reviews 85(4):935–956. https://doi.org/10.1111/j.1469-185X.2010.00141.x
- Nakagawa et al. (in prep) Quantifying between- and within-individual correlations and the degree of trait integration.