gllvmTMB fits generalised linear latent variable
models (GLLVMs) for unit × trait data — site × species,
individual × trait, species × trait, paper × item, or any other Unit ×
Trait formulation. The package is a TMB engine that supports
glmmTMB / brms-style covariance dispatch via
three covstruct modes expressed with four formula
keywords:
-
Paired decomposition —
latent + uniquetogether gives (shared low-rank plus per-trait unique). -
Marginal only —
indepalone gives (per-trait variance, no cross-trait shared structure). -
Full unstructured —
depalone gives free, parameterised via Cholesky.
Built on top: response distributions for binary / count / continuous / ordinal / hurdle data, sparse SPDE spatial random fields, phylogenetic random effects (sparse- representation for big trees), and multi-tier hierarchies — all turned on only when the biological question requires them. Each topical article in the side-bar walks through one such extension on a runnable example.
What a GLLVM estimates
A GLLVM models multivariate data: a matrix of responses with rows (units — sites, individuals, species, …) and columns (responses — species, traits, behaviours, items, …). It combines two familiar ideas:
- Like a generalised linear mixed model, each response gets its own distribution and link function so binary, count, and continuous responses are all on the same footing.
- Like factor analysis, the covariance among the responses is summarised by a small number of latent dimensions.
The expected value is linked to a linear predictor through , where
- are measured predictors with response-specific coefficients — the regression part of the model.
- is a length- vector of latent scores for unit , and is a length- loading vector for response . Stacking the loadings across responses gives a matrix whose -th row is .
In the Gaussian case (), the model can be written equivalently as
so the leftover — what’s not explained by the regression part and the shared latent dimensions — has variance specific to response . This is the response-specific unique variance (also called the “specific” variance in factor analysis). Stacking the such variances on the diagonal gives the diagonal matrix
The marginal covariance among responses (after measured predictors) then decomposes as
The two pieces are complementary:
-
is the
shared covariance among responses, induced through the
latent dimensions. Because
,
the shared component has reduced rank — covariance among many responses
is captured by a few latent dimensions instead of
free pairwise entries. In
gllvmTMB, is parameterised by the formula keywordlatent(0 + trait | unit, d = K). -
is the
diagonal of response-specific unique variances
(one entry per response, off-diagonal entries are zero by construction).
In
gllvmTMB, is parameterised byunique(0 + trait | unit).
A useful per-response summary is communality:
The numerator is the variance of response explained by the shared latent structure; the denominator is the total model-implied residual variance of response . So is the proportion of response ’s variance shared with the others. High means response is strongly integrated; low means response-specific variation dominates.
A fitted GLLVM therefore returns:
- Regression coefficients for measured predictors.
- Latent scores placing units in -dimensional latent space (model-based ordination).
- Loadings describing how strongly each response is associated with each latent dimension (biplot arrows).
- Residual covariance / correlation among responses.
- Communality per response.
Example: Individual × Trait morphology
We have n individuals, each measured on T
morphological traits across a few sessions. The biological question: do
the traits covary on a small number of latent axes (e.g. a body-size
axis, a shape axis)? With one observational level and modest repetition,
both the shared (latent()) and unique
(unique()) covariance components are identifiable. No
spatial or phylogenetic structure here.
Simulate n = 60 individuals × T = 5 traits
× 3 sessions with two true latent axes plus trait-specific unique
variance:
set.seed(1)
n_indiv <- 60
n_session <- 3
n_traits <- 5
trait_names <- paste0("trait_", seq_len(n_traits))
Lambda_true <- matrix(c( 1.0, 0.6, -0.4, 0.2, 0.5,
0.0, 0.8, 0.5, -0.3, 0.2),
nrow = n_traits, ncol = 2)
S_B_true <- c(0.20, 0.15, 0.30, 0.20, 0.25)
sigma_eps <- 0.20
alpha <- c(0, 0.5, -0.3, 0.2, 0.1)
# Stable per-individual trait values (between-individual rr + unique)
u <- matrix(rnorm(n_indiv * 2), n_indiv, 2)
between_ind <- matrix(alpha, n_indiv, n_traits, byrow = TRUE) +
u %*% t(Lambda_true) +
matrix(rnorm(n_indiv * n_traits,
sd = sqrt(S_B_true[col(matrix(0, n_indiv, n_traits))])),
n_indiv, n_traits)
# Three sessions per individual, each with measurement noise
df <- expand.grid(individual_id = seq_len(n_indiv),
session_id = seq_len(n_session),
trait_idx = seq_len(n_traits))
df$value <- between_ind[cbind(df$individual_id, df$trait_idx)] +
rnorm(nrow(df), sd = sigma_eps)
df$individual <- factor(df$individual_id)
df$trait <- factor(trait_names[df$trait_idx], levels = trait_names)
df$session <- factor(paste(df$individual_id, df$session_id, sep = "_"))
df <- df[, c("individual", "session", "trait", "value")]
head(df, 6)
#> individual session trait value
#> 1 1 1_1 trait_1 -0.5657110
#> 2 2 2_1 trait_1 0.6421943
#> 3 3 3_1 trait_1 -0.9446050
#> 4 4 4_1 trait_1 1.1630869
#> 5 5 5_1 trait_1 0.3986457
#> 6 6 6_1 trait_1 -0.1792850The DGP has three variance sources: shared low-rank
,
per-trait between-individual unique
,
and per-session noise
.
The fit below recovers the first two via latent() and
unique(); the per-session noise is absorbed into the
Gaussian residual variance
(one parameter per trait).
Fit a d = 2 reduced-rank GLLVM with
latent() + unique() at the individual
level — the shared low-rank structure plus trait-specific unique
variance:
fit <- gllvmTMB(
value ~ 0 + trait +
latent(0 + trait | individual, d = 2) +
unique(0 + trait | individual),
data = df,
unit = "individual"
)
fit
#> Stacked-trait gllvmTMB fit
#> Traits = 5, individual = 60
#> Covstructs: latent_unit, unique_unit
#> Fixed effects (b_fix): 5
#> log L = -359.627 convergence = 0
#> Note: Lambda_B identified up to rotation (use suggest_lambda_constraint() or rotate_loadings()).
#> Run gllvmTMB_diagnose(fit) for a full health check, or summary(fit) for parameter estimates.The recovered loadings (up to column rotation / sign — only is identifiable from the data):
Lambda_hat <- fit$report$Lambda_B
rownames(Lambda_hat) <- paste0("trait_", seq_len(n_traits))
colnames(Lambda_hat) <- paste0("LV", seq_len(2))
round(Lambda_hat, 3)
#> LV1 LV2
#> trait_1 0.884 0.000
#> trait_2 0.380 0.727
#> trait_3 -0.287 0.469
#> trait_4 0.285 -0.221
#> trait_5 0.420 0.031The implied trait covariance vs the truth (off-diagonal pattern is what should agree — diagonals include unique variance):
list(true_LL = round(Lambda_true %*% t(Lambda_true), 2),
fitted_LL = round(Lambda_hat %*% t(Lambda_hat), 2))
#> $true_LL
#> [,1] [,2] [,3] [,4] [,5]
#> [1,] 1.0 0.60 -0.40 0.20 0.50
#> [2,] 0.6 1.00 0.16 -0.12 0.46
#> [3,] -0.4 0.16 0.41 -0.23 -0.10
#> [4,] 0.2 -0.12 -0.23 0.13 0.04
#> [5,] 0.5 0.46 -0.10 0.04 0.29
#>
#> $fitted_LL
#> trait_1 trait_2 trait_3 trait_4 trait_5
#> trait_1 0.78 0.34 -0.25 0.25 0.37
#> trait_2 0.34 0.67 0.23 -0.05 0.18
#> trait_3 -0.25 0.23 0.30 -0.19 -0.11
#> trait_4 0.25 -0.05 -0.19 0.13 0.11
#> trait_5 0.37 0.18 -0.11 0.11 0.18Ordination of individuals on the two latent axes:
ord <- extract_ordination(fit, level = "unit")
plot(ord$scores[, 1], ord$scores[, 2], pch = 19, col = "steelblue",
xlab = "LV 1", ylab = "LV 2",
main = "Individuals on latent trait axes")
abline(h = 0, v = 0, lty = 2, col = "grey70")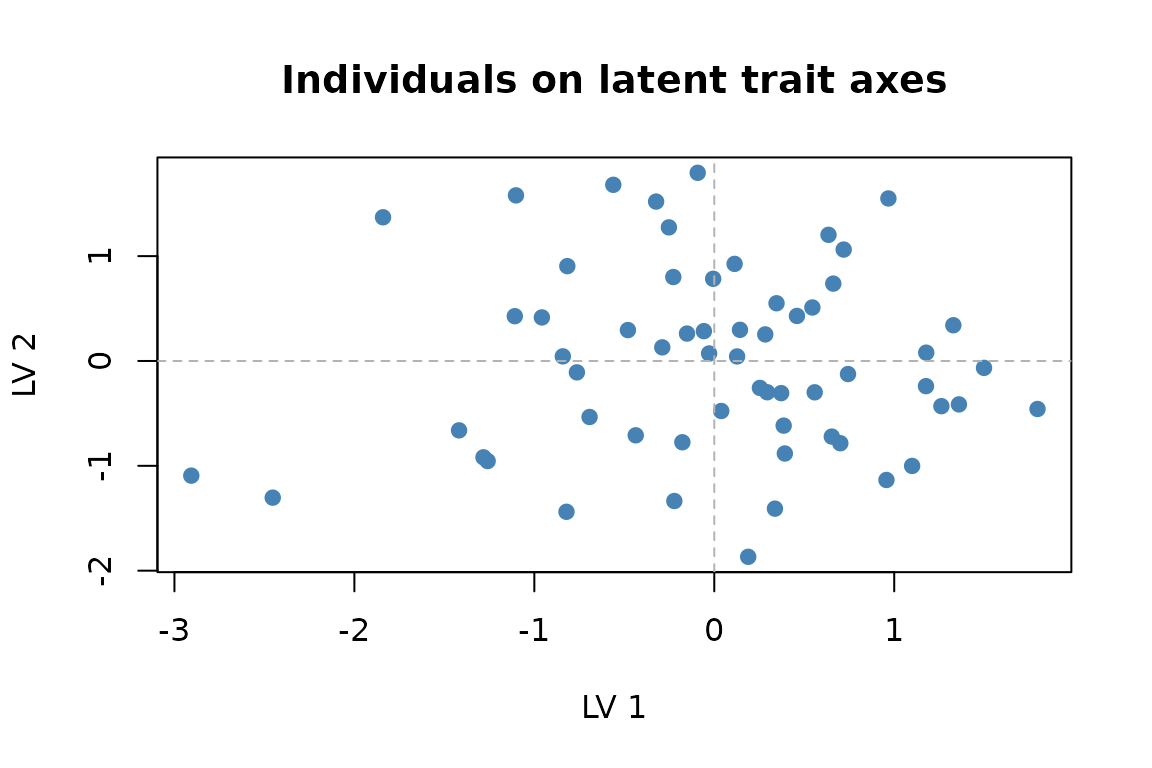
Individual scores on the two latent trait axes.
That’s the whole model — latent() for shared trait
covariance, unique() for trait-specific residual variance,
plus trait-specific intercepts.
Per-trait communality — the proportion of each trait’s variance shared with the others:
extract_communality(fit, "unit")
#> trait_1 trait_2 trait_3 trait_4 trait_5
#> 0.8275929 0.7731596 0.4767918 0.3807250 0.4533901Pairwise trait correlations with 95% Fisher-z Wald confidence
intervals. method = "fisher-z" is the default and keeps
bounds in [-1, 1]; method = "profile" and
method = "bootstrap" are slower alternatives, and
method = "wald" is kept as an alias of
"fisher-z":
extract_correlations(fit, tier = "unit")
#> tier trait_i trait_j correlation lower upper method
#> 1 B trait_1 trait_2 0.37022453 0.12836755 0.57051555 fisher-z
#> 2 B trait_1 trait_3 -0.32811832 -0.53727841 -0.08093708 fisher-z
#> 3 B trait_2 trait_3 0.31216578 0.06325509 0.52451403 fisher-z
#> 4 B trait_1 trait_4 0.44361843 0.21377450 0.62692297 fisher-z
#> 5 B trait_2 trait_4 -0.09622584 -0.34179899 0.16164860 fisher-z
#> 6 B trait_3 trait_4 -0.39849036 -0.59246553 -0.16084056 fisher-z
#> 7 B trait_1 trait_5 0.61087893 0.42248966 0.74867173 fisher-z
#> 8 B trait_2 trait_5 0.31206494 0.06314382 0.52443305 fisher-z
#> 9 B trait_3 trait_5 -0.21289593 -0.44287881 0.04337352 fisher-z
#> 10 B trait_4 trait_5 0.30863879 0.05936672 0.52167898 fisher-zThe matrix form, if you want to look at the correlation structure as a whole rather than per pair:
round(cov2cor(extract_Sigma(fit, level = "unit")$Sigma), 2)
#> trait_1 trait_2 trait_3 trait_4 trait_5
#> trait_1 1.00 0.37 -0.33 0.44 0.61
#> trait_2 0.37 1.00 0.31 -0.10 0.31
#> trait_3 -0.33 0.31 1.00 -0.40 -0.21
#> trait_4 0.44 -0.10 -0.40 1.00 0.31
#> trait_5 0.61 0.31 -0.21 0.31 1.00Acknowledgements
gllvmTMB is a standalone TMB-based package. Its design
borrows ideas from several earlier projects, whose corresponding GPL-3
code is incorporated and whose authors are credited in
DESCRIPTION (as copyright holders) and listed in
inst/COPYRIGHTS:
- sdmTMB (Anderson, Ward, English, Barnett, Thorson, et al.): sparse SPDE / GMRF spatial machinery.
-
glmmTMB
(Brooks, Bolker, Kristensen, Magnusson, Skaug, Nielsen, Berg, van
Bentham, et al.):
rr/propto/equalto/diagcovariance dispatch. The reduced-rankrr()machinery in particular is the contribution of McGillycuddy, Popovic, Bolker & Warton (2025) J. Stat. Softw. 112(1). -
GALAMM
(Sørensen): mixed-response (
gfam) and confirmatory fixed-loading (lambda_constraint) patterns. -
MCMCglmm
(Hadfield): sparse
phylogeny representation used by
phylo_latent(). - TMB (Kristensen et al.) and fmesher (Lindgren): autodiff and finite-element mesh.
