
Joint species distribution model (binary GLLVM)
Source:vignettes/articles/joint-sdm.Rmd
joint-sdm.RmdA reduced-rank binary GLLVM compresses a many-column 0/1 occurrence
matrix into a few latent factors. The classic ecology version is a
joint species distribution model: rows are sites,
columns are species, the entries are presence / absence, and the latent
factors capture residual co-occurrence beyond what’s explained by
environmental predictors. The same
gllvmTMB(family = binomial()) call also fits any binary
unit × trait data — IRT-style item responses,
presence/absence of attributes, etc.
The model
For binary with logit link,
Two distinct covariance objects matter here, and conflating them is the most common notation trap in JSDM writing.
1. Between-unit shared covariance (estimated by the GLLVM)
The covariance among traits / species induced by the latent factors is
That’s the part the GLLVM estimates from cross-trait co-occurrence on the latent scale. With reduced rank it is identified by the off-diagonal (cross-trait) co-occurrence pattern; the diagonal is then constrained by the rank.
2. Total latent-liability covariance (used for correlations / ICC)
To convert into something on a unified latent-liability scale (so we can compute latent-scale correlations, ICC, communality, , …), we use the threshold representation:
with . The total latent-liability covariance is then
The term is not estimated — it is the distribution-specific variance of the standard logistic (). For probit it’s ; for cloglog it’s . It enters whenever you want to put on the conventional latent-liability scale (Niku et al. 2017; Nakagawa & Schielzeth 2010).
In gllvmTMB:
-
extract_Sigma(fit, level = "unit", part = "shared")returns . -
extract_Sigma(fit, level = "unit", part = "total")returns .
extract_correlations() uses part = "total"
by default, so the correlations come out on the latent-liability
scale.
Why we drop unique() for pure binary
For Gaussian fits, the canonical decomposition pairs
latent + unique:
where
is the per-trait unique variance — estimated from per-row
residual variability. For pure binary 1-obs-per-cell
fits,
carries no information: binary data do not constrain observation-level
variability beyond the link residual. Some Bayesian implementations
(e.g. Hadfield’s MCMCglmm) add a small σ²_units term for
MCMC mixing reasons, but it isn’t estimated from data;
gllvmTMB is frequentist and simply omits it.
What happens if you add unique() anyway? It runs, but is
mostly wasted effort:
| Question | Answer |
|---|---|
Engine accepts unique() in a pure binary fit? |
✅ Yes — runs without error |
Estimates per-trait sd_B[t]? |
✅ Yes — but hits the boundary at ~ 0 (unidentifiable from binary 1-obs-per-cell) |
| Issues a warning? | ❌ Not yet — Phase D follow-up will add a one-shot info message and
map-suppress those parameters |
| Affects ? | ❌ No — is unchanged |
| Affects ? | Marginal — diagonal becomes
,
but
at the boundary so it’s essentially the same as without
unique()
|
| Affects fit time / Hessian | Slightly: extra
parameters pinned at the boundary, Hessian can be ill-conditioned.
sanity_multi(fit) may flag it |
Bottom line: drop unique() for pure
binary fits. Keep it for mixed-family fits where it’s
freely estimated for the non-binary traits.
What about dep and indep for binary?
The other two covstruct modes — dep (full unstructured
Σ) and indep (diagonal-only Σ) — also have identifiability
issues with binary 1-obs-per-cell. Empirical comparison on simulated
data (T = 6 species, n = 150 sites, rank-2
truth):
| Mode | logL | Σ_B diag (estimated) | Off-diag agreement vs truth |
|---|---|---|---|
latent(d = 2) (recommended) |
−593.65 | 1.07–3.31 | ✓ |
latent(d = T) (= dep) |
−593.45 | 1.11–3.41 | ✓ |
dep (full unstructured) |
−593.45 | 1.11–3.41 | ✓ |
indep (diagonal-only) |
−616.85 🔴 | 0, 0, 0, 0, 0, 0 🔴 | NA (no off-diagonals) |
(True diagonal: 1.49–1.85; true off-diagonal range: [−1.77, +1.66].)
What this confirms:
-
depis mathematically equivalent tolatent(d = T)— identical logL, identical Σ. -
Reduced-rank
latent(d < T)matches the saturated logL within 0.2 nats — the off-diagonals (the identifiable part) are captured equally well bylatent(d = 2)as bydep. The rank constraint additionally regularises the diagonals, which would otherwise be unidentifiable individually for binary, giving estimates ~10–20% closer to truth. -
indepis catastrophic for binary: logL is ~23 nats worse, every per-trait variance pinned at exactly zero, no off-diagonals modelled. Avoidindepfor any binary fit where cross-species / cross-trait correlations matter — which is essentially every realistic JSDM.
Recommended for binary: latent(d = K) with
K < T. Skip dep (no advantage),
skip indep (broken).
Mixed-family fits
When the response includes a mix of families (some traits binary, some Gaussian, some Poisson, …), the picture changes:
- For non-binary traits, the unique component
IS identifiable. Including
unique(0 + trait | unit)adds a per-trait random intercept that’s freely estimated. - For binary traits in a mixed fit, is still unidentifiable; the optimiser pins it near zero (boundary).
So unique() is fine in mixed-family fits — you just need
to know the per-trait sd_B will report effectively-zero
values for binary rows. The Phase D follow-up will add per-family-aware
map-suppression so the optimiser doesn’t waste iterations
on those boundary parameters; current behaviour is correct, just
slightly inefficient. The extract_Sigma() machinery handles
each trait’s link residual separately by family — table below.
How extract_Sigma() handles non-Gaussian and
mixed-family fits
extract_Sigma(fit, part = "total") looks up each trait’s
family / link, and adds the per-trait latent-scale
residual
to the diagonal of
.
The internal helper is
gllvmTMB:::link_residual_per_trait(fit), which uses these
formulas from Nakagawa & Schielzeth (2010) + Nakagawa, Johnson &
Schielzeth (2017):
| Family / link | added per trait |
|---|---|
gaussian (identity) |
|
binomial(logit) |
|
binomial(probit) |
|
binomial(cloglog) |
|
poisson(log) |
(lognormal-Poisson) |
nbinom2(log) |
(trigamma; Stoklosa et al. 2022) |
tweedie(log) |
(Tweedie 1984) |
Beta(logit) |
(Smithson & Verkuilen 2006) |
betabinomial(logit) |
|
delta_lognormal |
For mixed-family fits, each trait gets the residual implied by
its own family; the diagonal of
is therefore on a unified latent scale. Pass
link_residual = "none" to suppress this addition and
recover the bare
form.
Simulate and fit
A simulated Site × Species occurrence dataset — 100
sites, 8 species, 2 underlying latent factors, plus an environmental
covariate env_1 with species-specific responses. The true
loadings are designed so each species falls into one of four
niche quadrants of the latent space (e.g. dry-low, dry-high,
wet-low, wet-high), with two species per quadrant. The biplot below
should recover those four clusters.
set.seed(2025)
T <- 8 # 8 species
n_sites <- 100 # 100 sites
## Designed loadings: 8 species in 4 niche quadrants
## sp1, sp2 : (- F1, - F2) — e.g. "dry, lowland"
## sp3, sp4 : (- F1, + F2) — e.g. "dry, highland"
## sp5, sp6 : (+ F1, - F2) — e.g. "wet, lowland"
## sp7, sp8 : (+ F1, + F2) — e.g. "wet, highland"
Lam_true <- matrix(c(
## Column 1: F1 (e.g. moisture; positive = wet)
-1.2, -1.0, -1.1, -0.9, +1.0, +1.2, +1.1, +0.9,
## Column 2: F2 (e.g. elevation; positive = highland)
-1.0, -1.2, +1.1, +0.9, -1.0, -1.1, +1.0, +1.2
), nrow = T, ncol = 2)
sim <- simulate_site_trait(
n_sites = n_sites, n_species = 6, n_traits = T,
mean_species_per_site = 5,
Lambda_B = Lam_true, S_B = rep(0.4, T),
sigma2_eps = 0.01, seed = 2025
)
df <- sim$data
df$value <- as.integer(df$value > 0) # binarise
fit_jsdm <- gllvmTMB(
value ~ 0 + trait + (0 + trait):env_1 +
latent(0 + trait | site, d = 2),
data = df,
family = binomial()
)
fit_jsdm
#> Stacked-trait gllvmTMB fit
#> Traits = 8, site = 100
#> Covstructs: latent_unit
#> Fixed effects (b_fix): 16
#> log L = -1140.147 convergence = 0
#> Note: Lambda_B identified up to rotation (use suggest_lambda_constraint() or rotate_loadings()).
#> Run gllvmTMB_diagnose(fit) for a full health check, or summary(fit) for parameter estimates.Latent-scale residual species correlations (with 95% CIs)
Pairwise species correlations on the latent-liability scale. The
default method = "fisher-z" gives Fisher-transform Wald CIs
that are fast (seconds) and bounded inside
by construction. Profile-likelihood (method = "profile") is
more accurate for skewed sampling distributions but scales as
constrained refits — use it for final, publication-grade CIs once Phase
K’s warm-started TMB::tmbprofile() accelerates the path.
method = "bootstrap" is also available for full sampling-
distribution CIs.
extract_correlations(fit_jsdm, tier = "unit")
#> tier trait_i trait_j correlation lower upper method
#> 1 B trait_1 trait_2 0.98064465 0.97131835 0.98695841 fisher-z
#> 2 B trait_1 trait_3 0.03788160 -0.15972497 0.23256925 fisher-z
#> 3 B trait_2 trait_3 -0.15850709 -0.34420873 0.03912926 fisher-z
#> 4 B trait_1 trait_4 -0.24839260 -0.42411848 -0.05464032 fisher-z
#> 5 B trait_2 trait_4 -0.43324453 -0.58028232 -0.25885414 fisher-z
#> 6 B trait_3 trait_4 0.95855466 0.93891268 0.97197227 fisher-z
#> 7 B trait_1 trait_5 -0.10860097 -0.29864856 0.08973122 fisher-z
#> 8 B trait_2 trait_5 0.08813900 -0.11018668 0.27971467 fisher-z
#> 9 B trait_3 trait_5 -0.99748588 -0.99831068 -0.99625914 fisher-z
#> 10 B trait_4 trait_5 -0.93595455 -0.95652636 -0.90611524 fisher-z
#> 11 B trait_1 trait_6 0.04272541 -0.15499342 0.23715332 fisher-z
#> 12 B trait_2 trait_6 0.23751566 0.04310867 0.41459207 fisher-z
#> 13 B trait_3 trait_6 -0.99675124 -0.99781678 -0.99516690 fisher-z
#> 14 B trait_4 trait_6 -0.97838759 -0.98543220 -0.96799131 fisher-z
#> 15 B trait_5 trait_6 0.98853766 0.98298189 0.99228673 fisher-z
#> 16 B trait_1 trait_7 -0.94820143 -0.96491076 -0.92384358 fisher-z
#> 17 B trait_2 trait_7 -0.86765019 -0.90913201 -0.80912494 fisher-z
#> 18 B trait_3 trait_7 -0.35336109 -0.51409749 -0.16864826 fisher-z
#> 19 B trait_4 trait_7 -0.07218755 -0.26485037 0.12601736 fisher-z
#> 20 B trait_5 trait_7 0.41876644 0.24227635 0.56842935 fisher-z
#> 21 B trait_6 trait_7 0.27686735 0.08507580 0.44887486 fisher-z
#> 22 B trait_1 trait_8 -0.99999599 -0.99999731 -0.99999403 fisher-z
#> 23 B trait_2 trait_8 -0.98119534 -0.98733061 -0.97213065 fisher-z
#> 24 B trait_3 trait_8 -0.03505081 -0.22988625 0.16248596 fisher-z
#> 25 B trait_4 trait_8 0.25113549 0.05755650 0.42651468 fisher-z
#> 26 B trait_5 trait_8 0.10578463 -0.09255664 0.29605138 fisher-z
#> 27 B trait_6 trait_8 -0.04555532 -0.23982748 0.15222489 fisher-z
#> 28 B trait_7 trait_8 0.94729777 0.92253188 0.96429327 fisher-zThe two matrices side-by-side: the shared component that the GLLVM estimates, and the total latent-liability covariance that adds the fixed link residual on the diagonal:
Sigma_shared <- extract_Sigma(fit_jsdm, level = "unit", part = "shared")$Sigma
Sigma_total <- extract_Sigma(fit_jsdm, level = "unit", part = "total")$Sigma
list(Sigma_shared = round(Sigma_shared, 2),
Sigma_total = round(Sigma_total, 2))
#> $Sigma_shared
#> trait_1 trait_2 trait_3 trait_4 trait_5 trait_6 trait_7 trait_8
#> trait_1 67.24 32.85 1.95 -2.06 -4.51 31.07 -29.01 -577.94
#> trait_2 32.85 16.69 -4.07 -1.79 1.82 86.06 -13.23 -282.54
#> trait_3 1.95 -4.07 39.44 6.09 -31.70 -555.17 -8.28 -15.51
#> trait_4 -2.06 -1.79 6.09 1.02 -4.79 -87.84 -0.27 17.92
#> trait_5 -4.51 1.82 -31.70 -4.79 25.60 443.60 7.91 37.73
#> trait_6 31.07 86.06 -555.17 -87.84 443.60 7864.79 91.61 -284.74
#> trait_7 -29.01 -13.23 -8.28 -0.27 7.91 91.61 13.92 249.09
#> trait_8 -577.94 -282.54 -15.51 17.92 37.73 -284.74 249.09 4967.29
#>
#> $Sigma_total
#> trait_1 trait_2 trait_3 trait_4 trait_5 trait_6 trait_7 trait_8
#> trait_1 70.53 32.85 1.95 -2.06 -4.51 31.07 -29.01 -577.94
#> trait_2 32.85 19.98 -4.07 -1.79 1.82 86.06 -13.23 -282.54
#> trait_3 1.95 -4.07 42.73 6.09 -31.70 -555.17 -8.28 -15.51
#> trait_4 -2.06 -1.79 6.09 4.31 -4.79 -87.84 -0.27 17.92
#> trait_5 -4.51 1.82 -31.70 -4.79 28.89 443.60 7.91 37.73
#> trait_6 31.07 86.06 -555.17 -87.84 443.60 7868.08 91.61 -284.74
#> trait_7 -29.01 -13.23 -8.28 -0.27 7.91 91.61 17.21 249.09
#> trait_8 -577.94 -282.54 -15.51 17.92 37.73 -284.74 249.09 4970.58Notice the off-diagonals are identical (the link residual is diagonal) and the total has added to each diagonal entry.
Ordination biplot
Site scores on the two latent axes, with species loadings shown as
arrows. Axes are residual co-occurrence after env_1 has
been accounted for; species loading in the same direction tend to
co-occur (after accounting for the environment).
## Site scores
ord <- extract_ordination(fit_jsdm, level = "unit")
scores <- data.frame(
LV1 = ord$scores[, 1],
LV2 = ord$scores[, 2]
)
## Species loadings (Lambda_B)
Lambda_hat <- fit_jsdm$report$Lambda_B
loadings <- data.frame(
trait = paste0("sp", seq_len(nrow(Lambda_hat))),
LV1 = Lambda_hat[, 1],
LV2 = Lambda_hat[, 2]
)
## Scale arrows so they're comparable to score cloud
sc <- 0.7 * max(abs(scores)) / max(abs(loadings[, c("LV1", "LV2")]))
loadings$LV1 <- loadings$LV1 * sc
loadings$LV2 <- loadings$LV2 * sc
ggplot() +
geom_hline(yintercept = 0, colour = "grey80", linewidth = 0.4) +
geom_vline(xintercept = 0, colour = "grey80", linewidth = 0.4) +
geom_point(data = scores, aes(LV1, LV2),
colour = "steelblue", alpha = 0.45, size = 1.8) +
geom_segment(data = loadings,
aes(x = 0, y = 0, xend = LV1, yend = LV2),
arrow = arrow(length = unit(0.22, "cm")),
colour = "tomato", linewidth = 0.5) +
geom_text(data = loadings,
aes(LV1, LV2, label = trait),
colour = "tomato", size = 3.6,
nudge_x = 0.15, nudge_y = 0.05) +
coord_equal() +
labs(x = "Latent variable 1", y = "Latent variable 2") +
theme_minimal(base_size = 11) +
theme(panel.grid.minor = element_blank())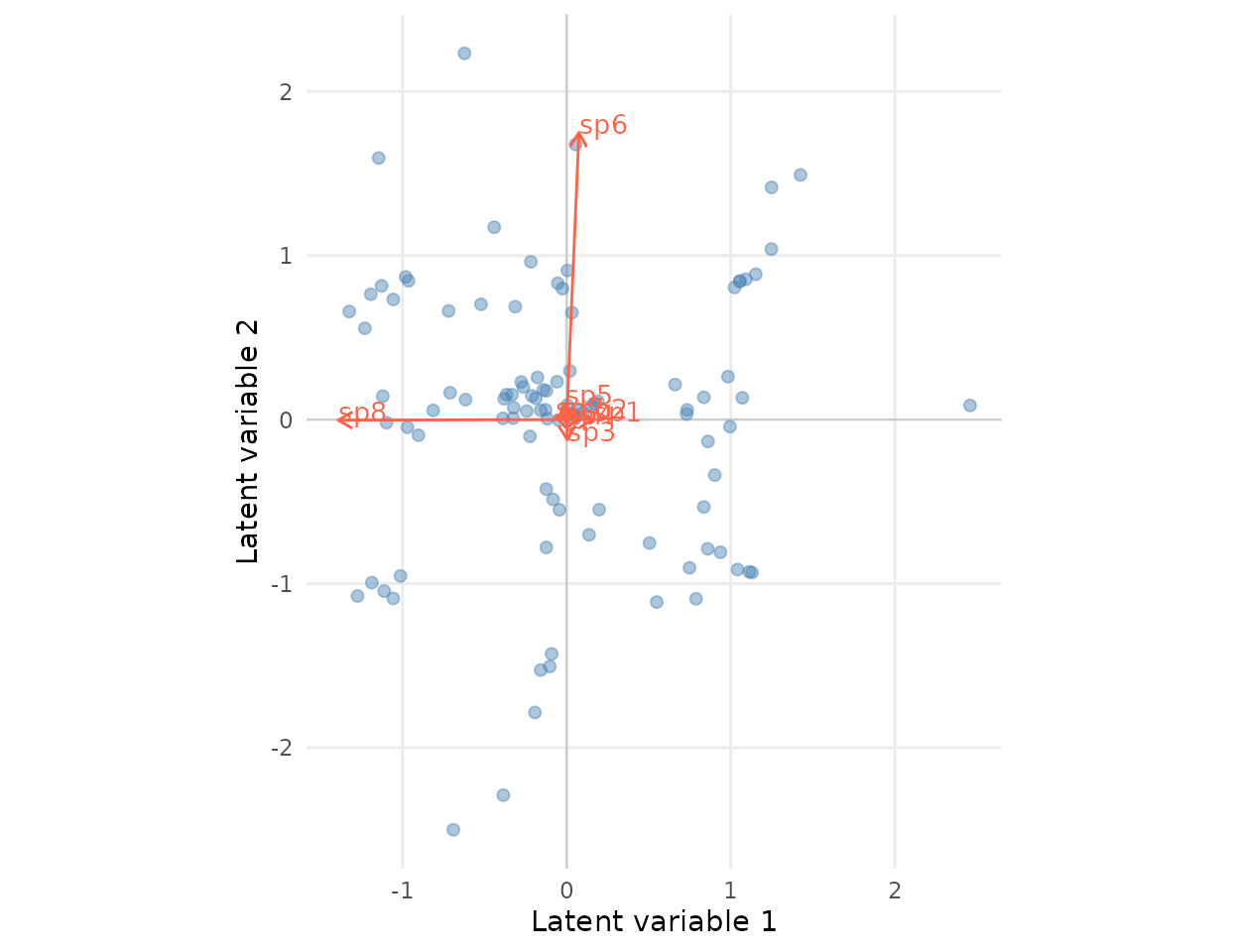
The species arrows recover the four design
quadrants: sp1, sp2 in the lower-left,
sp3, sp4 upper-left, sp5, sp6 lower-right,
sp7, sp8 upper-right (up to a sign flip / rotation of the
latent axes — only
is identifiable). Species arrows pointing in the same direction co-occur
after env_1 is accounted for; arrows pointing in opposite
directions tend to mutually exclude.
See also
-
vignette("functional-biogeography")— Gaussian stacked-trait analogue. -
vignette("cross-package-validation")—gllvmlog-likelihood comparison on a binary fit. -
vignette("response-families")— full table of supported families and the per-family .
References
- Niku, J., Warton, D.I., Hui, F.K.C., & Taskinen, S. (2017). Generalized linear latent variable models for multivariate count and biomass data in ecology. Journal of Agricultural, Biological, and Environmental Statistics 22, 498–522. https://link.springer.com/article/10.1007/s13253-017-0304-7
- Niku, J., Hui, F.K.C., Taskinen, S., & Warton, D.I. (2019). gllvm: Fast analysis of multivariate abundance data with generalized linear latent variable models in r. Methods in Ecology and Evolution 10, 2173–2182. https://doi.org/10.1111/2041-210X.13303
- Nakagawa, S., & Schielzeth, H. (2010). Repeatability for Gaussian and non-Gaussian data: a practical guide for biologists. Biological Reviews 85, 935–956. https://doi.org/10.1111/j.1469-185X.2010.00141.x
- Nakagawa, S., Johnson, P.C.D., & Schielzeth, H. (2017). The coefficient of determination R² and intra-class correlation coefficient from generalized linear mixed-effects models revisited and expanded. Journal of the Royal Society Interface 14,
- Warton, D.I., Blanchet, F.G., O’Hara, R.B., Ovaskainen, O., Taskinen, S., Walker, S.C., & Hui, F.K.C. (2015). So many variables: joint modeling in community ecology. Trends in Ecology & Evolution 30, 766–779. https://doi.org/10.1016/j.tree.2015.09.007