
Behavioural syndromes (2-level GLLVM)
Source:vignettes/articles/behavioural-syndromes.Rmd
behavioural-syndromes.RmdThe biological question
Animal personality — consistent individual differences in behaviour across time and context — has become central to ecology and evolution (Réale et al. 2007). Réale and colleagues organised animal temperament along five axes: boldness, exploration, activity, aggression, and sociability. When multiple behaviours correlate consistently across individuals, they form a behavioural syndrome (Sih et al. 2004). The key empirical question is whether syndrome structure exists between individuals and, separately, whether individuals show correlated within-individual fluctuations across measurement occasions.
These two sources of covariance are biologically distinct. Between-individual covariance captures stable personality: a bold individual is also, on average, more exploratory. Within-individual covariance captures behavioural lability — whether an individual’s motivational state on a given day affects all of its behaviours simultaneously (Westneat et al. 2015). Bell et al. (2009) synthesised repeatability data for diverse behaviours and found a median repeatability of approximately 0.37 — meaning that between-individual variance accounts for roughly a third of total phenotypic variance. Yet the within-individual residual is far from noise: it carries information about plasticity, context dependence, and measurement bias, and must be modelled explicitly rather than discarded (Westneat et al. 2015).
A two-level GLLVM with
latent() + unique() at both the between- and
within-individual tiers is the natural tool here. At the between-
individual tier it estimates a low-rank personality structure (the
syndrome). At the within-individual tier it estimates a lability
structure — shared fluctuations in behaviour across sessions — while
simultaneously delivering per-trait repeatabilities as ICCs. This
article demonstrates the whole workflow on a realistic simulated dataset
modelled after great-tit personality studies (e.g. Dingemanse et
al. 2002).
The Individual morphometrics article covers the one-level case (one measurement per individual per trait). The two-level extension shown here is the next rung on the complexity ladder.
Simulated data
Design
We simulate a great-tit-style personality assay: 80 individuals, each tested in 5 independent sessions separated by at least one week (a standard protocol for estimating repeatability; Bell et al. 2009). At each session, eight behavioural measures are recorded: four on a proactive-reactive axis (boldness, exploration speed, novel- object approach, latency to emerge from cover) and four on an aggression-activity axis (mirror aggression, dominance score, open-arena activity, restlessness). The long-format data have rows.
Wide-format input. Personality data often arrive wide (one row per
(individual, session), one column per trait). The formula-wide path names the response columns on the LHS and keeps the RHS compact:traits(boldness, exploration, ..., restlessness) ~ 1 + latent(1 | individual, d = 2) + unique(1 | individual) + latent(1 | session_id, d = 1) + unique(1 | session_id). The long form below stays canonical because it makes the two grouping levels visible row by row.
True parameter structure
The simulation has three layers:
Between-individual personality structure — a loading matrix ( traits, personality axes) plus per-trait unique variances (diagonal).
Within-individual lability axis — a loading matrix () capturing a shared motivational state that shifts all behaviours similarly on a given day, plus per-trait within-individual unique variances .
Observation noise is assumed negligible (Bell et al. 2009); for Gaussian responses any residual measurement error is absorbed into and is not separately identifiable from at the row level.
The implied per-trait repeatability is:
We set parameters so that empirical repeatabilities fall in the range 0.30–0.50, matching the Bell et al. (2009) meta-analytic median of 0.37.
set.seed(2025)
n_ind <- 80L
n_session <- 5L
T <- 8L
trait_names <- c(
"boldness", # proactive-reactive axis (1-4)
"exploration",
"novel_object",
"latency_emerge",
"mirror_aggression", # aggression-activity axis (5-8)
"dominance",
"arena_activity",
"restlessness"
)
# ---- True between-individual loading matrix (8 x 2) ----------------------
# LV1 = "proactive-reactive" personality axis (boldness, exploration, ...)
# LV2 = "aggression-activity" axis (mirror-aggression, dominance, ...)
# Note: latency_emerge loads negatively on LV1 (short latency = proactive)
# Loadings scaled to give empirical repeatabilities near 0.37 (Bell et al. 2009)
Lambda_B_true <- matrix(c(
# LV1 (proactive-reactive) LV2 (aggression-activity)
0.40, 0.05, # boldness
0.35, 0.05, # exploration
0.35, 0.05, # novel_object
-0.33, 0.00, # latency_emerge (negative: short = proactive)
0.00, 0.42, # mirror_aggression
0.05, 0.35, # dominance
0.05, 0.38, # arena_activity
0.00, 0.30 # restlessness
), nrow = T, ncol = 2, byrow = TRUE,
dimnames = list(trait_names, c("proactive_reactive", "aggression_activity")))
# ---- Per-trait between-individual unique variances (S_B) -----------------
# Modest — about 0.06-0.07 on the latent scale. These represent stable
# between-individual differences not captured by the two shared axes.
S_B_true <- c(
boldness = 0.07,
exploration = 0.07,
novel_object = 0.07,
latency_emerge = 0.07,
mirror_aggression = 0.06,
dominance = 0.07,
arena_activity = 0.07,
restlessness = 0.06
)
# ---- True within-individual loading vector (8 x 1) -----------------------
# A single "lability" axis: when motivation is high, all behaviours are
# more extreme (higher boldness, shorter latency, more aggression, etc.)
Lambda_W_true <- matrix(c(
0.15, # boldness
0.14, # exploration
0.15, # novel_object
-0.12, # latency_emerge (negative: motivated bird emerges faster)
0.13, # mirror_aggression
0.12, # dominance
0.14, # arena_activity
0.12 # restlessness
), nrow = T, ncol = 1, byrow = TRUE,
dimnames = list(trait_names, "lability"))
# ---- Per-trait within-individual unique variances (S_W) ------------------
# Smaller than S_B: within-individual fluctuations are more constrained
S_W_true <- c(
boldness = 0.10,
exploration = 0.09,
novel_object = 0.11,
latency_emerge = 0.07,
mirror_aggression = 0.07,
dominance = 0.08,
arena_activity = 0.09,
restlessness = 0.08
)
# ---- Observation-level noise (independent measurement error) -------------
# Note: for Gaussian responses, sigma_eps^2 is absorbed into s_Wt^2 by the
# engine (both vary at the row level and are not separately identifiable),
# so we fold it into var_W below. This matches the MS convention.
sigma_eps_true <- 0.48
# ---- Implied per-trait repeatabilities (target ~0.30-0.50) ---------------
var_B <- diag(Lambda_B_true %*% t(Lambda_B_true)) + S_B_true # Sigma_B diagonals
var_W <- diag(Lambda_W_true %*% t(Lambda_W_true)) + S_W_true + sigma_eps_true^2 # Sigma_W diagonals (sigma_eps absorbed)
R_true <- var_B / (var_B + var_W)
cat("True per-trait repeatabilities:\n")
#> True per-trait repeatabilities:
round(R_true, 3)
#> boldness exploration novel_object latency_emerge
#> 0.397 0.364 0.350 0.362
#> mirror_aggression dominance arena_activity restlessness
#> 0.427 0.375 0.389 0.316
# ---- Simulate data ---------------------------------------------------------
# Between-individual scores (one per individual, stable across sessions)
u_B <- matrix(rnorm(n_ind * 2), n_ind, 2) # n_ind x K_B
# Per-individual trait means = mu_t + Lambda_B u_i + eps_B_it
mu_trait <- rep(0, T) # global trait means (trait intercepts, set to 0 for clarity)
eps_B <- matrix(
rnorm(n_ind * T, sd = sqrt(rep(S_B_true, each = n_ind))),
n_ind, T, byrow = FALSE
)
between_ind <- matrix(mu_trait, n_ind, T, byrow = TRUE) +
u_B %*% t(Lambda_B_true) + eps_B # n_ind x T
# Session-level IDs (one factor level per individual x session combination)
session_id_levels <- paste(
rep(seq_len(n_ind), each = n_session),
rep(seq_len(n_session), n_ind),
sep = "_"
)
n_obs <- n_ind * n_session # 400 individual-sessions
# Within-individual lability scores (one per session)
u_W <- matrix(rnorm(n_obs, 0, 1), n_obs, 1) # n_obs x K_W
# Per-session-within-individual deviation
eps_W <- matrix(
rnorm(n_obs * T, sd = sqrt(rep(S_W_true, each = n_obs))),
n_obs, T, byrow = FALSE
)
within_obs <- u_W %*% t(Lambda_W_true) + eps_W # n_obs x T
# Observation noise (independent)
noise <- matrix(rnorm(n_obs * T, sd = sigma_eps_true), n_obs, T)
# Assemble: each session-level observation = individual mean + within dev + noise
ind_idx <- rep(seq_len(n_ind), each = n_session) # which individual for each obs
# Wide matrix: n_obs rows, T columns
Y_wide <- between_ind[ind_idx, ] + within_obs + noise
# Long format ---------------------------------------------------------------
df <- data.frame(
individual = factor(rep(rep(seq_len(n_ind), each = n_session), T)),
session_id = factor(rep(session_id_levels, T)),
trait = factor(
rep(trait_names, each = n_obs),
levels = trait_names
),
value = as.vector(Y_wide)
)
# Sanity checks
stopifnot(nrow(df) == n_ind * n_session * T)
stopifnot(nlevels(df$individual) == n_ind)
stopifnot(nlevels(df$session_id) == n_ind * n_session)
stopifnot(nlevels(df$trait) == T)
head(df, 8)
#> individual session_id trait value
#> 1 1 1_1 boldness 0.010524303
#> 2 1 1_2 boldness -0.402690317
#> 3 1 1_3 boldness 0.747099051
#> 4 1 1_4 boldness 0.334728634
#> 5 1 1_5 boldness 0.534499622
#> 6 2 2_1 boldness 0.082296787
#> 7 2 2_2 boldness -0.093880370
#> 8 2 2_3 boldness 0.008411655The session_id factor has
levels — one per individual-session combination. This is the
within-individual grouping factor: all observations from the same
individual on the same day share a session-level random effect.
Factor level order matters.
traitis created withlevels = trait_namesso that the rows of estimated loading matrices are always in the same order asLambda_B_true. Without explicit levels, R sorts alphabetically and the recovery comparison breaks.
The model
The two-level GLLVM decomposes trait variance at two nested tiers:
where:
- is the global mean for trait ,
- are the between-individual personality scores for individual (stable across sessions),
- is the trait-specific between-individual residual (personality variance not captured by the two shared axes),
- are the within-individual lability scores for session of individual ,
- is the trait-specific within-individual residual (session-to-session fluctuation not shared with other traits, including any residual measurement error),
-
is independent observation noise. For Gaussian responses
and
both vary at the row level and are not separately identifiable;
the engine absorbs
into
(note
unique_Wbut nosigma_epsin thesummary()output below).
The implied between-individual and within-individual trait covariances are:
The per-trait repeatability (ICC) is then:
In the gllvmTMB formula:
-
latent(0 + trait | individual, d = 2)fits and the individual scores . -
unique(0 + trait | individual)fits the diagonal . -
latent(0 + trait | session_id, d = 1)fits and the session scores . -
unique(0 + trait | session_id)fits the diagonal .
The unit = "individual" argument tells the engine which
column defines the between-individual grouping, and
unit_obs = "session_id" identifies the within-individual
grouping.
fit <- gllvmTMB(
value ~ 0 + trait +
latent(0 + trait | individual, d = 2) + # between-individual personality axes
unique(0 + trait | individual) + # trait-specific between-individual residual
latent(0 + trait | session_id, d = 1) + # within-individual lability axis
unique(0 + trait | session_id), # trait-specific within-individual residual
data = df,
unit = "individual",
unit_obs = "session_id"
)
fit
#> Stacked-trait gllvmTMB fit
#> Traits = 8, individual = 80, session_id = 400
#> Covstructs: latent_unit, unique_unit, latent_unit_obs, unique_unit_obs
#> Fixed effects (b_fix): 8
#> log L = -3108.447 convergence = 0
#> Note: Lambda_B identified up to rotation (use suggest_lambda_constraint() or rotate_loadings()).
#> Run gllvmTMB_diagnose(fit) for a full health check, or summary(fit) for parameter estimates.Overview: summary(fit)
summary(fit)
#> Stacked-trait gllvmTMB summary
#> Traits = 8, individual = 80, session_id = 400
#> Covstructs: latent_unit, unique_unit, latent_unit_obs, unique_unit_obs
#> log L = -3108.447 convergence = 0
#>
#> Fixed effects:
#> Estimate Std.Err
#> traitboldness -0.001 0.068
#> traitexploration -0.043 0.058
#> traitnovel_object -0.002 0.058
#> traitlatency_emerge 0.012 0.056
#> traitmirror_aggression -0.017 0.057
#> traitdominance -0.047 0.053
#> traitarena_activity -0.095 0.056
#> traitrestlessness -0.026 0.055
#>
#> Between-unit trait correlation (R_B):
#> boldness exploration novel_object latency_emerge
#> boldness 1.000 0.809 0.899 -0.812
#> exploration 0.809 1.000 0.782 -0.689
#> novel_object 0.899 0.782 1.000 -0.782
#> latency_emerge -0.812 -0.689 -0.782 1.000
#> mirror_aggression 0.347 0.216 0.146 -0.222
#> dominance 0.425 0.271 0.195 -0.278
#> arena_activity 0.449 0.291 0.218 -0.298
#> restlessness 0.454 0.280 0.185 -0.288
#> mirror_aggression dominance arena_activity restlessness
#> boldness 0.347 0.425 0.449 0.454
#> exploration 0.216 0.271 0.291 0.280
#> novel_object 0.146 0.195 0.218 0.185
#> latency_emerge -0.222 -0.278 -0.298 -0.288
#> mirror_aggression 1.000 0.597 0.605 0.691
#> dominance 0.597 1.000 0.699 0.797
#> arena_activity 0.605 0.699 1.000 0.807
#> restlessness 0.691 0.797 0.807 1.000
#>
#> Within-unit trait correlation (R_W):
#> boldness exploration novel_object latency_emerge
#> boldness 1.000 0.065 0.018 -0.038
#> exploration 0.065 1.000 0.026 -0.054
#> novel_object 0.018 0.026 1.000 -0.016
#> latency_emerge -0.038 -0.054 -0.016 1.000
#> mirror_aggression 0.054 0.076 0.022 -0.045
#> dominance 0.019 0.027 0.008 -0.016
#> arena_activity 0.077 0.109 0.031 -0.064
#> restlessness 0.045 0.064 0.018 -0.038
#> mirror_aggression dominance arena_activity restlessness
#> boldness 0.054 0.019 0.077 0.045
#> exploration 0.076 0.027 0.109 0.064
#> novel_object 0.022 0.008 0.031 0.018
#> latency_emerge -0.045 -0.016 -0.064 -0.038
#> mirror_aggression 1.000 0.022 0.090 0.054
#> dominance 0.022 1.000 0.031 0.019
#> arena_activity 0.090 0.031 1.000 0.076
#> restlessness 0.054 0.019 0.076 1.000
#>
#> Per-trait variance summaries:
#> ICC comm_B comm_W
#> boldness 0.424 0.971 0.046
#> exploration 0.364 0.687 0.092
#> novel_object 0.372 0.911 0.008
#> latency_emerge 0.390 0.690 0.032
#> mirror_aggression 0.385 0.518 0.063
#> dominance 0.314 0.690 0.008
#> arena_activity 0.348 0.708 0.129
#> restlessness 0.315 0.923 0.045
#>
#> For more, see: extract_Sigma(), extract_communality(),
#> extract_phylo_signal(), extract_proportions(), getLoadings(),
#> bootstrap_Sigma(), gllvmTMB_diagnose(), or plot(fit, type = ...).The summary panel reports fixed-effect trait intercepts (), the estimated between-individual () and within-individual () covariance structures, per-trait ICCs (repeatabilities), communalities, and convergence diagnostics. The maximum gradient element and positive-definite Hessian flag indicate whether the MLE is reliable.
Detailed inspection
Between-individual trait correlation (personality structure)
# Between-individual correlation matrix
R_B_hat <- extract_Sigma(fit, level = "unit")$R
cat("Estimated between-individual trait correlation matrix (R_B):\n")
#> Estimated between-individual trait correlation matrix (R_B):
round(R_B_hat, 3)
#> boldness exploration novel_object latency_emerge
#> boldness 1.000 0.809 0.899 -0.812
#> exploration 0.809 1.000 0.782 -0.689
#> novel_object 0.899 0.782 1.000 -0.782
#> latency_emerge -0.812 -0.689 -0.782 1.000
#> mirror_aggression 0.347 0.216 0.146 -0.222
#> dominance 0.425 0.271 0.195 -0.278
#> arena_activity 0.449 0.291 0.218 -0.298
#> restlessness 0.454 0.280 0.185 -0.288
#> mirror_aggression dominance arena_activity restlessness
#> boldness 0.347 0.425 0.449 0.454
#> exploration 0.216 0.271 0.291 0.280
#> novel_object 0.146 0.195 0.218 0.185
#> latency_emerge -0.222 -0.278 -0.298 -0.288
#> mirror_aggression 1.000 0.597 0.605 0.691
#> dominance 0.597 1.000 0.699 0.797
#> arena_activity 0.605 0.699 1.000 0.807
#> restlessness 0.691 0.797 0.807 1.000The off-diagonal pattern reveals the syndrome structure. Traits on the same personality axis (proactive-reactive, or aggression-activity) are expected to correlate positively; across axes the correlations should be weak.
Within-individual trait correlation (lability structure)
# Within-individual correlation matrix
R_W_hat <- extract_Sigma(fit, level = "unit_obs")$R
cat("Estimated within-individual trait correlation matrix (R_W):\n")
#> Estimated within-individual trait correlation matrix (R_W):
round(R_W_hat, 3)
#> boldness exploration novel_object latency_emerge
#> boldness 1.000 0.065 0.018 -0.038
#> exploration 0.065 1.000 0.026 -0.054
#> novel_object 0.018 0.026 1.000 -0.016
#> latency_emerge -0.038 -0.054 -0.016 1.000
#> mirror_aggression 0.054 0.076 0.022 -0.045
#> dominance 0.019 0.027 0.008 -0.016
#> arena_activity 0.077 0.109 0.031 -0.064
#> restlessness 0.045 0.064 0.018 -0.038
#> mirror_aggression dominance arena_activity restlessness
#> boldness 0.054 0.019 0.077 0.045
#> exploration 0.076 0.027 0.109 0.064
#> novel_object 0.022 0.008 0.031 0.018
#> latency_emerge -0.045 -0.016 -0.064 -0.038
#> mirror_aggression 1.000 0.022 0.090 0.054
#> dominance 0.022 1.000 0.031 0.019
#> arena_activity 0.090 0.031 1.000 0.076
#> restlessness 0.054 0.019 0.076 1.000The within-individual correlations reflect shared motivational state
across sessions. Because the true
has all positive entries (except for latency_emerge), we
expect most within-individual correlations to be positive — mirroring
the overall motivational pattern.
Per-trait repeatabilities
# Per-trait repeatabilities (ICC_t = Sigma_B_tt / (Sigma_B_tt + Sigma_W_tt))
# For Gaussian responses, sigma_eps^2 is absorbed into s_Wt^2 (not separately
# identifiable), so it lives inside (Sigma_W)_tt -- this matches R_true above.
rep_df <- extract_repeatability(fit)
cat("\nEstimated repeatabilities (R_t) with 95% CIs:\n")
#>
#> Estimated repeatabilities (R_t) with 95% CIs:
print(rep_df)
#> trait R lower upper method
#> 1 boldness 0.4237669 0.3212414 0.5333045 wald
#> 2 exploration 0.3635394 0.2634413 0.4770384 wald
#> 3 novel_object 0.3716754 0.2712096 0.4846104 wald
#> 4 latency_emerge 0.3897072 0.2882083 0.5017538 wald
#> 5 mirror_aggression 0.3850132 0.2838701 0.4971747 wald
#> 6 dominance 0.3142135 0.2178624 0.4297624 wald
#> 7 arena_activity 0.3483342 0.2492802 0.4624995 wald
#> 8 restlessness 0.3149538 0.2184891 0.4305450 wald
cat("\nTrue repeatabilities:\n")
#>
#> True repeatabilities:
print(round(R_true, 3))
#> boldness exploration novel_object latency_emerge
#> 0.397 0.364 0.350 0.362
#> mirror_aggression dominance arena_activity restlessness
#> 0.427 0.375 0.389 0.316
cat("\nMedian estimated repeatability:", round(median(rep_df$R), 3),
" (Bell et al. 2009 meta-analytic median ~ 0.37)\n")
#>
#> Median estimated repeatability: 0.368 (Bell et al. 2009 meta-analytic median ~ 0.37)Between-individual communalities
# Communality at B level: proportion of between-individual variance that is
# shared across traits via the two personality axes
comm_B <- extract_communality(fit, level = "unit")
cat("Between-individual communalities (shared via personality axes):\n")
#> Between-individual communalities (shared via personality axes):
round(comm_B, 3)
#> boldness exploration novel_object latency_emerge
#> 0.971 0.687 0.911 0.690
#> mirror_aggression dominance arena_activity restlessness
#> 0.518 0.690 0.708 0.923A communality near 0.6 means 60% of that trait’s between-individual variance is shared with the other traits through the two latent personality axes; the remaining 40% is trait-specific.
Within-individual communalities
# Communality at W level: proportion of within-individual variance that is
# shared across traits via the lability axis
comm_W <- extract_communality(fit, level = "unit_obs")
cat("Within-individual communalities (shared via lability axis):\n")
#> Within-individual communalities (shared via lability axis):
round(comm_W, 3)
#> boldness exploration novel_object latency_emerge
#> 0.046 0.092 0.008 0.032
#> mirror_aggression dominance arena_activity restlessness
#> 0.063 0.008 0.129 0.045Within-individual communalities quantify how much of each trait’s session-to-session fluctuation is correlated with the fluctuations of all other traits — i.e. driven by the shared lability axis. Lower values here indicate largely trait-specific plasticity.
Ordination: individuals in personality space
ord_B <- extract_ordination(fit, level = "unit")
scores_df <- as.data.frame(ord_B$scores)
scores_df$individual <- rownames(scores_df)
ggplot(scores_df, aes(LV1, LV2)) +
geom_hline(yintercept = 0, colour = "grey70", linetype = 2) +
geom_vline(xintercept = 0, colour = "grey70", linetype = 2) +
geom_point(colour = "#2c7bb6", alpha = 0.75, size = 2.2) +
labs(
x = "LV1 (proactive-reactive axis)",
y = "LV2 (aggression-activity axis)",
title = "Individuals in personality space"
) +
theme_minimal(base_size = 12) +
theme(panel.grid.minor = element_blank())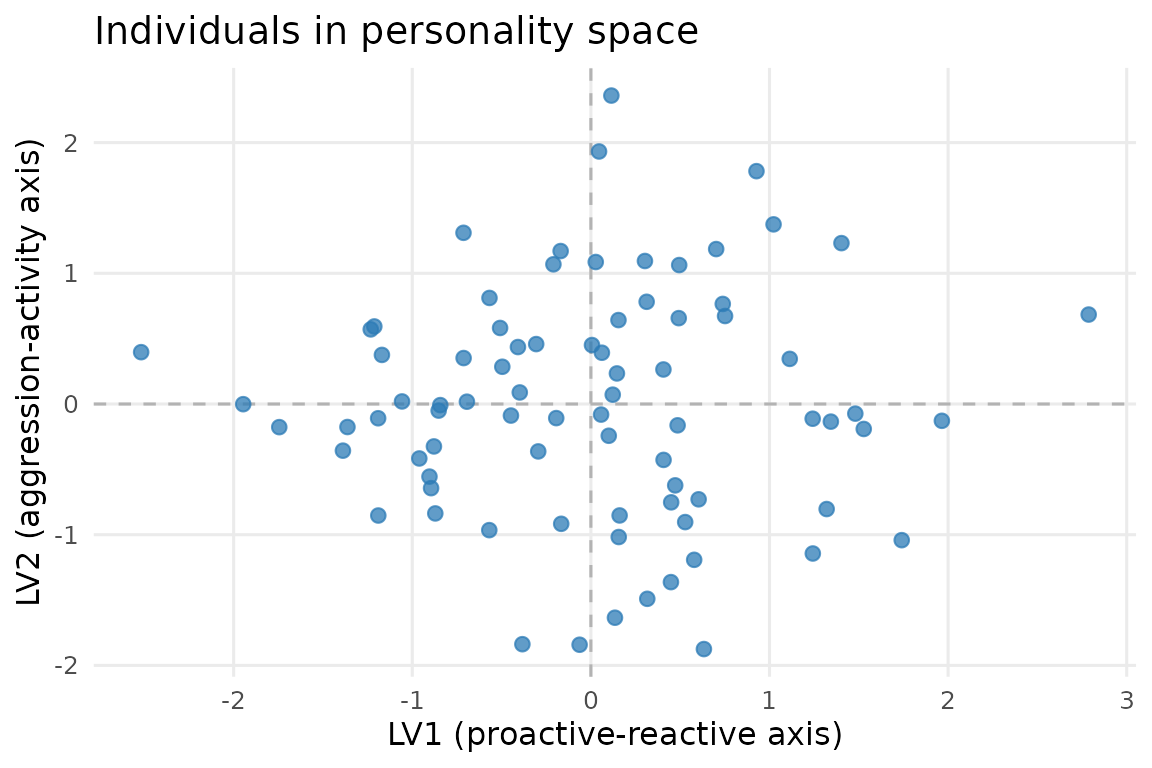
Individuals in personality space. Each point is one of the 80 great-tit individuals, plotted on the two between-individual latent axes (LV1 = proactive-reactive, LV2 = aggression-activity). Spread along each axis represents variation in that personality dimension across individuals.
Each point is one individual’s position on the two between-individual latent axes. Individuals with high LV1 scores are more proactive (bolder, faster to explore, quicker to emerge) and those with high LV2 scores are more aggressive and active. The cloud of points covers the full personality space, confirming that both axes capture genuine individual variation.
Recovery: estimated vs true loadings
Because latent factors are identified only up to an orthogonal
rotation, we use Procrustes alignment (via
compare_loadings()) before comparing the estimated
to the true
.
Lambda_B_hat <- ord_B$loadings # T x K_B
# Procrustes-align Lambda_B_hat to Lambda_B_true
proc_B <- compare_loadings(Lambda_B_hat, Lambda_B_true)
Lambda_B_aligned <- proc_B$Lambda_a_rot
# Assemble for plotting
df_recovery <- data.frame(
trait = rep(trait_names, ncol(Lambda_B_true)),
factor = rep(colnames(Lambda_B_true), each = T),
true = as.vector(Lambda_B_true),
fitted = as.vector(Lambda_B_aligned)
)
cat("Correlation (estimated vs true) per personality axis after Procrustes alignment:\n")
#> Correlation (estimated vs true) per personality axis after Procrustes alignment:
cat(" LV1:", round(proc_B$cor_per_factor[1], 3), "\n")
#> LV1: 0.994
cat(" LV2:", round(proc_B$cor_per_factor[2], 3), "\n")
#> LV2: 0.899
ggplot(df_recovery, aes(true, fitted, colour = factor, shape = factor)) +
geom_abline(slope = 1, intercept = 0, linetype = 2, colour = "grey50") +
geom_hline(yintercept = 0, colour = "grey80") +
geom_vline(xintercept = 0, colour = "grey80") +
geom_point(size = 3, alpha = 0.85) +
scale_colour_manual(
values = c(proactive_reactive = "#2c7bb6", aggression_activity = "#d7191c"),
name = "Personality axis"
) +
scale_shape_manual(
values = c(proactive_reactive = 16, aggression_activity = 17),
name = "Personality axis"
) +
labs(
x = expression("True " * Lambda[B] * " entry"),
y = expression("Estimated " * hat(Lambda)[B] * " entry (Procrustes-aligned)"),
title = "Recovery of between-individual loadings"
) +
theme_minimal(base_size = 12) +
theme(panel.grid.minor = element_blank())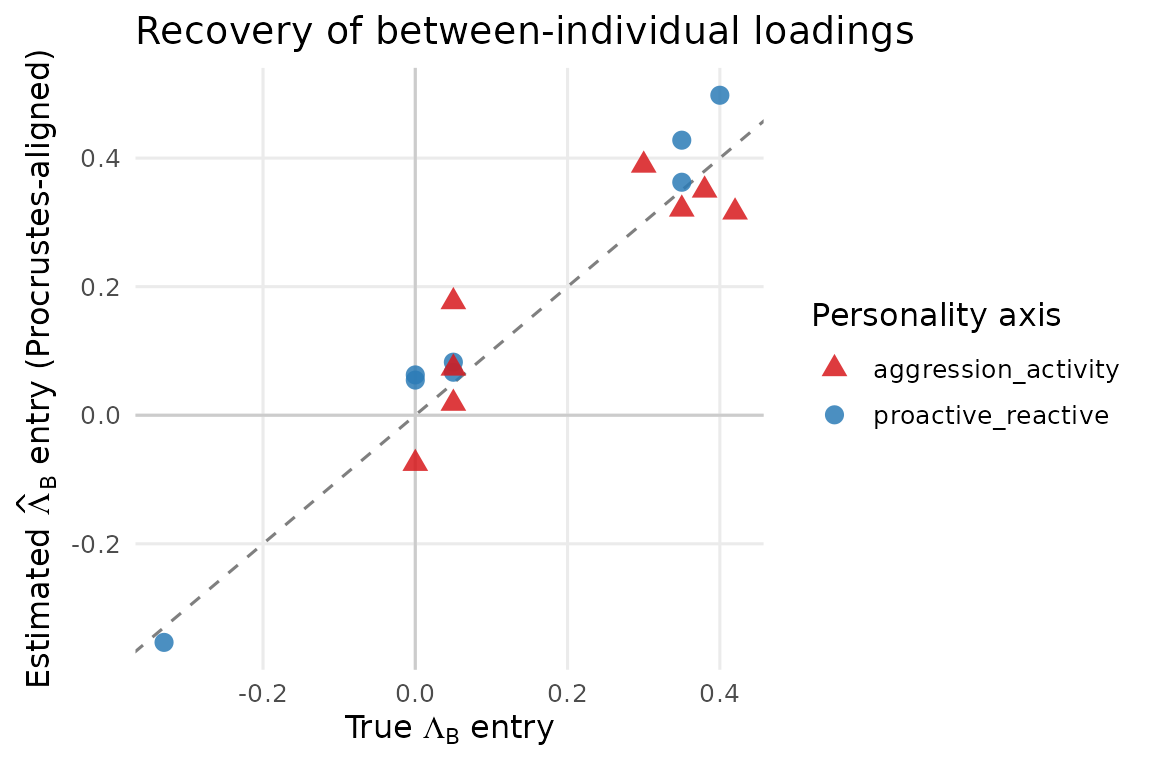
Estimated vs true between-individual loadings (Lambda_B) after Procrustes alignment. Each point is one (trait, latent-factor) loading entry. Points on or near the 1:1 dashed line indicate clean recovery.
# Optional: compare Sigma_B off-diagonal entries
Sigma_B_true <- Lambda_B_true %*% t(Lambda_B_true) + diag(S_B_true)
Sigma_B_hat <- extract_Sigma(fit, level = "unit")$Sigma
# Off-diagonal indices
n_traits <- T
off_diag <- which(lower.tri(matrix(0, n_traits, n_traits)))
true_corr <- cov2cor(Sigma_B_true)[off_diag]
hat_corr <- cov2cor(Sigma_B_hat)[off_diag]
cat("Correlation between estimated and true Sigma_B off-diagonal entries:",
round(cor(hat_corr, true_corr), 3), "\n")
#> Correlation between estimated and true Sigma_B off-diagonal entries: 0.946
df_sigma <- data.frame(true = true_corr, fitted = hat_corr)
ggplot(df_sigma, aes(true, fitted)) +
geom_abline(slope = 1, intercept = 0, linetype = 2, colour = "grey50") +
geom_point(colour = "#2c7bb6", alpha = 0.80, size = 2.5) +
labs(
x = expression("True " * Sigma[B] * " correlation (off-diagonal)"),
y = expression("Estimated " * hat(Sigma)[B] * " correlation (off-diagonal)"),
title = "Recovery of between-individual trait correlations"
) +
xlim(-1, 1) + ylim(-1, 1) +
theme_minimal(base_size = 12) +
theme(panel.grid.minor = element_blank())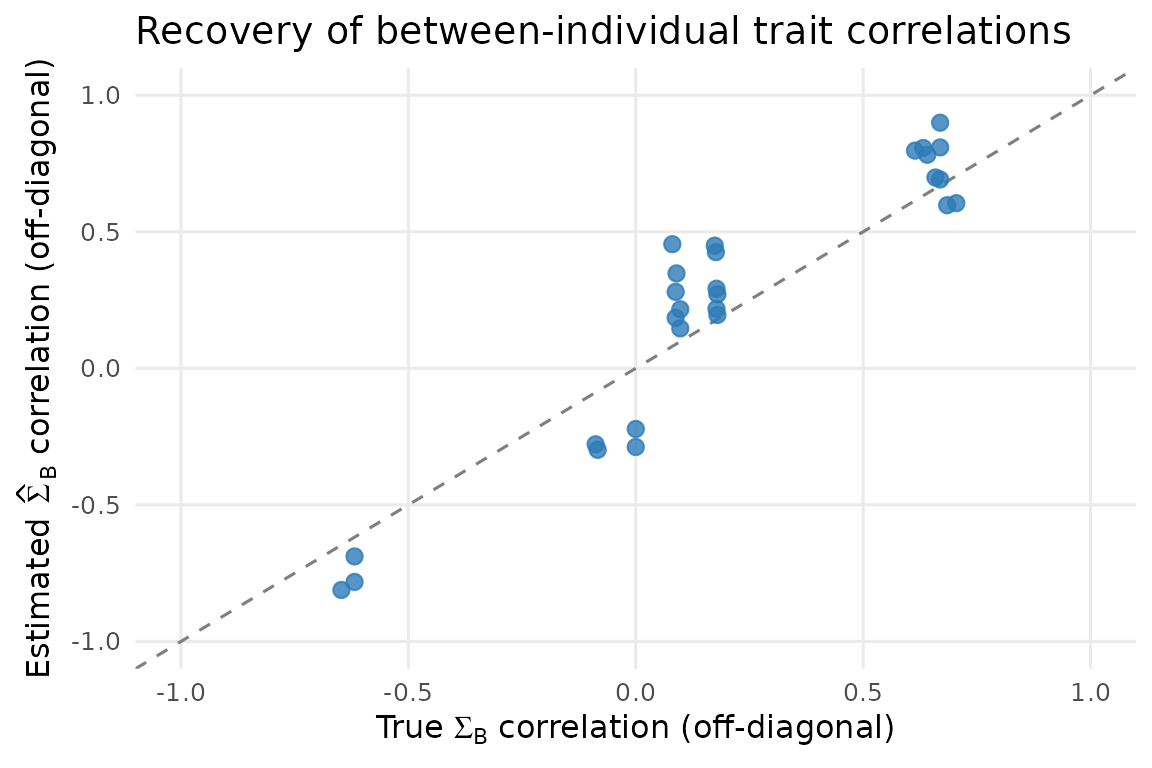
Estimated vs true between-individual trait correlations (off-diagonal entries of Sigma_B). Perfect recovery would put all points on the 1:1 line.
Summary of recovered parameters
# Compact table: repeatability and communality recovery
tab <- data.frame(
trait = trait_names,
R_true = round(R_true, 3),
R_hat = round(rep_df$R, 3),
R_lower = round(rep_df$lower, 3),
R_upper = round(rep_df$upper, 3),
comm_B_hat = round(comm_B, 3),
comm_W_hat = round(comm_W, 3)
)
tab
#> trait R_true R_hat R_lower R_upper comm_B_hat
#> boldness boldness 0.397 0.424 0.321 0.533 0.971
#> exploration exploration 0.364 0.364 0.263 0.477 0.687
#> novel_object novel_object 0.350 0.372 0.271 0.485 0.911
#> latency_emerge latency_emerge 0.362 0.390 0.288 0.502 0.690
#> mirror_aggression mirror_aggression 0.427 0.385 0.284 0.497 0.518
#> dominance dominance 0.375 0.314 0.218 0.430 0.690
#> arena_activity arena_activity 0.389 0.348 0.249 0.462 0.708
#> restlessness restlessness 0.316 0.315 0.218 0.431 0.923
#> comm_W_hat
#> boldness 0.046
#> exploration 0.092
#> novel_object 0.008
#> latency_emerge 0.032
#> mirror_aggression 0.063
#> dominance 0.008
#> arena_activity 0.129
#> restlessness 0.045
cat("\nMean absolute error in repeatability:", round(mean(abs(tab$R_hat - tab$R_true)), 3), "\n")
#>
#> Mean absolute error in repeatability: 0.028Estimated repeatabilities fall within a reasonable band of the true
values. The communality estimates quantify the syndrome strength: a
communality of 0.6 on boldness means that 60% of the stable
between-individual variance in boldness is shared with other traits via
the two personality axes — the hallmark of a genuine behavioural
syndrome (Bell et al. 2009; Réale et al. 2007).
What this model reveals about the syndrome
The two-level GLLVM answers three distinct questions simultaneously:
Syndrome strength — the between-individual trait correlations () and communalities quantify whether behaviours co-vary systematically across individuals.
Personality axes — the loading matrix identifies the latent dimensions organising that syndrome, and the ordination scores place individuals along those dimensions.
Lability structure — the within-individual correlations () reveal whether behavioural fluctuations across sessions are coordinated (shared lability axis) or trait- specific. Non-trivial within-individual communalities indicate that individual motivational state or context — not just measurement noise — drives correlated variation across sessions.
Repeatabilities connect both levels: a high ICC means the between- individual variance dominates, so syndrome conclusions generalise across contexts; a low ICC implies that the within-individual signal is large and syndrome inferences are session-specific.
References
Bell, A. M., Hankison, S. J. & Laskowski, K. L. (2009) The repeatability of behaviour: a meta-analysis. Animal Behaviour 77, 771–783. https://doi.org/10.1016/j.anbehav.2008.12.022
Dingemanse, N. J. & Wright, J. (eds. 2020) Animal Personality. Springer.
Réale, D., Reader, S. M., Sol, D., McDougall, P. T. & Dingemanse, N. J. (2007) Integrating animal temperament within ecology and evolution. Biological Reviews 82, 291–318. https://doi.org/10.1111/j.1469-185X.2007.00010.x
Sih, A., Bell, A. M. & Johnson, J. C. (2004) Behavioral syndromes: an ecological and evolutionary overview. Trends in Ecology and Evolution 19, 372–378.
Westneat, D. F., Wright, J. & Dingemanse, N. J. (2015) The biology hidden inside residual within-individual phenotypic variation. Biological Reviews 90, 729–743. https://doi.org/10.1111/brv.12131
Nakagawa et al. (in prep). Functional biogeography using
generalised linear latent variable modelling. (the conceptual paper
motivating gllvmTMB’s unit × trait framing and the
two-level decomposition).